 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aradu.BVL00 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Fabales; Fabaceae; Papilionoideae; Dalbergieae; Arachis
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 496aa MW: 54840.7 Da PI: 6.8795 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 54 | 3.7e-17 | 83 | 127 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+eE+e++++a k++G g W+ I +++g ++t+ q++s+ qk+
Aradu.BVL00 83 REKWTEEEHEKFLEALKLYGRG-WRQIEEHIG-TKTAVQIRSHAQKF 127
789*******************.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.14E-16 | 77 | 130 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 21.33 | 78 | 132 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.6E-10 | 80 | 130 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.0E-16 | 81 | 130 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 3.6E-13 | 82 | 130 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.3E-14 | 83 | 126 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 1.97E-11 | 85 | 128 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 496 aa Download sequence Send to blast |
MIQFSNMLVG LVPVKLRVFT RTGSDFAPME MQDQIEGTKL SKSGTASDCH SNDGSQSQSP 60 DISSVVNNHA PKVRKPYTIT KQREKWTEEE HEKFLEALKL YGRGWRQIEE HIGTKTAVQI 120 RSHAQKFFSK VVRDSDGSAE SSIHPIDIPP PRPKRKPLHP YPRKLANSFK RHSIPNEPEK 180 SPSPNLSSAE KDSQSPTSVL SAFGSEAFGS AFSEQTNRCL SPNSCTTDIQ SINLSPIEKE 240 NDCMTSKPSK EEDKTPLASA PLSISSQPYL HMKSEKLELC SKQTKCINND VVNVLPTTIK 300 LFGRTVSMVG NQESINTEEE CVKLCITRSD EVDDVENQLN TQLSLGMCNG SWQVMPKEAI 360 VKEDLGLGES APEASLSWSS WFPGGPAAYF GSSNTVVNPM PLHPSLKVRT REEEGSCTGS 420 NGESVNDVEI QGKIQGRFLD TVDSRCQQTS HQEEVVVPQK SPRGFVPYKR CLAERDASSL 480 IVALEERKGQ RARVCS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in phytochrome A-mediated cotyledon opening. Controlled by the central oscillator mediated by LHY and CCA1. Part of a regulatory circadian feedback loop. Regulates its own expression. {ECO:0000269|PubMed:14523250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00146 | DAP | Transfer from AT1G18330 | Download |
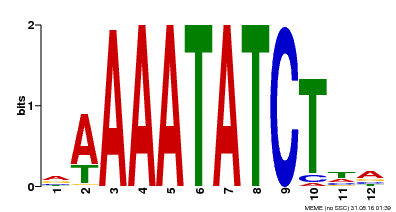
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Aradu.BVL00 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Circadian-regulation. Peak of transcript abundance near subjective dawn. Up-regulated transiently by red light and far red light. {ECO:0000269|PubMed:14523250, ECO:0000269|PubMed:19805390}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015967014.1 | 0.0 | protein REVEILLE 7 isoform X1 | ||||
| Swissprot | B3H5A8 | 5e-66 | RVE7_ARATH; Protein REVEILLE 7 | ||||
| TrEMBL | A0A445D127 | 0.0 | A0A445D127_ARAHY; Uncharacterized protein | ||||
| STRING | GLYMA16G34340.1 | 0.0 | (Glycine max) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF7313 | 32 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G18330.2 | 2e-61 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



