 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aco023605.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Bromeliaceae; Ananas
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 289aa MW: 32022.5 Da PI: 7.1505 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.3 | 9.3e-19 | 44 | 94 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s++ GVr+++ +grWvAeI+d + ++ r +lg+f taeeAa+a+++a+ l+g
Aco023605.1 44 SKFVGVRQRP-SGRWVAEIKDTT---QKIRMWLGTFETAEEAARAYDEAACLLRG 94
79********.7*********32...35**********************99987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 2.02E-31 | 44 | 101 | No hit | No description |
| SuperFamily | SSF54171 | 2.22E-19 | 44 | 102 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 2.9E-13 | 44 | 94 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 6.8E-28 | 45 | 102 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.559 | 45 | 102 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 5.0E-36 | 45 | 108 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.0E-7 | 46 | 57 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.0E-7 | 68 | 84 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0035865 | Biological Process | cellular response to potassium ion | ||||
| GO:0048528 | Biological Process | post-embryonic root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 289 aa Download sequence Send to blast |
MELHFQQQQQ QQQQQQQQQQ QYYLGTAASN KPNNKLKGKS RSSSKFVGVR QRPSGRWVAE 60 IKDTTQKIRM WLGTFETAEE AARAYDEAAC LLRGSNTRTN FVAHPSSPNS PLASRIRSLL 120 DHKKLKKSNT SPSNSTSITN TSISTSSTAA TTCSSTAATS ETMTNGTVST GATPCGDMKH 180 DTQLSTDHEV FRPYFINGSE DARLGSAHAD QSWAFEAGFD RLAVFDGYGE LANLINPEVS 240 EFERMRVERQ ISASLYAMNG VQEYFETVLD PCDLLWDLPP LCHLFCRS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 8e-14 | 37 | 101 | 6 | 71 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in the symbiotic nodule signaling pathway in response to rhizobial stimulation. Functions as a transcriptional regulator required for root infection by symbiotic rhizobia, infection thread (IT) formation, and nodule development. May coordinate these processes (PubMed:28503742, PubMed:28028038, PubMed:28068194). Functions downstream of the CCAMK-CYCLOPS complex (PubMed:28503742, PubMed:28028038). Probably not involved in arbuscular mycorrhizal (AM) symbiosis (PubMed:28068194). {ECO:0000269|PubMed:28028038, ECO:0000269|PubMed:28068194, ECO:0000269|PubMed:28503742}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
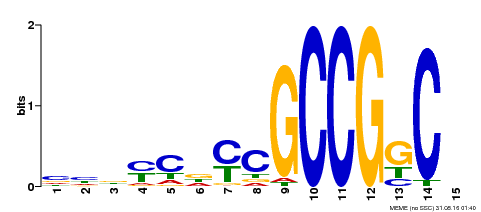
| Motif ID | Method | Source | Motif file |
| MP00521 | DAP | Transfer from AT5G19790 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced in nodules 7 days after rhizobial inoculation. {ECO:0000269|PubMed:28028038, ECO:0000269|PubMed:28503742}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020096589.1 | 0.0 | ethylene-responsive transcription factor RAP2-11-like | ||||
| Swissprot | A0A1X9PY88 | 5e-78 | ERN1_LOTJA; Ethylene-responsive transcription factor ERN1 | ||||
| TrEMBL | A0A199UJ14 | 0.0 | A0A199UJ14_ANACO; Ethylene-responsive transcription factor RAP2-11 | ||||
| STRING | XP_008804361.1 | 1e-100 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3900 | 32 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G19790.1 | 6e-28 | related to AP2 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aco023605.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



