 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aco015166.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Bromeliaceae; Ananas
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 830aa MW: 91347.5 Da PI: 6.1221 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.9 | 4.1e-19 | 18 | 76 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
Aco015166.1 18 KYVRYTPEQVEALERLYHECPKPSSMRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 76
5679*****************************************************97 PP
| |||||||
| 2 | START | 173.7 | 1.1e-54 | 162 | 370 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEEEEX CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlmvae 97
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s+++ g a+ra+g+v ++a v e+l+d + W++++++++++ v+++g g+++l +++
Aco015166.1 162 IAEETLAEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCAGVAARACGLVGIEPA-KVVEFLKDQPSWSRDCRSVDVVSVLTTGnnGTVELLYMQ 258
7899******************************************************.5556666666***************************** PP
XTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHH CS
START 98 lqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvksglae 191
l+a+++l+p Rd + Ry+ l++g++v++++S++++q p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ +++++++r+l++s+++
Aco015166.1 259 LYAPTTLAPaRDLWLMRYTSVLEDGSLVVCERSLNNTQGGPTmpsVQPFVRAEMLPSGYLIRPCDDGGSVIHIVYHMDLEPWSIPEVIRPLYESSTML 356
****************************************99888899************************************************** PP
HHHHHHHHTXXXXX CS
START 192 gaktwvatlqrqce 205
+++t++ +l+++++
Aco015166.1 357 AQRTTMTALRHLRQ 370
*********99875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.726 | 13 | 77 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 2.7E-16 | 15 | 81 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 3.94E-17 | 16 | 78 | No hit | No description |
| SuperFamily | SSF46689 | 4.71E-17 | 17 | 79 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 8.9E-17 | 18 | 76 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 9.7E-19 | 20 | 76 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 6.31E-5 | 70 | 109 | No hit | No description |
| PROSITE profile | PS50848 | 26.245 | 152 | 366 | IPR002913 | START domain |
| CDD | cd08875 | 1.75E-75 | 156 | 372 | No hit | No description |
| SMART | SM00234 | 2.2E-39 | 161 | 371 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 9.06E-38 | 161 | 372 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 2.9E-22 | 161 | 367 | IPR023393 | START-like domain |
| Pfam | PF01852 | 5.6E-52 | 162 | 370 | IPR002913 | START domain |
| Pfam | PF08670 | 1.2E-48 | 686 | 828 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 830 aa Download sequence Send to blast |
MMVVSSCKEG KGGMDPGKYV RYTPEQVEAL ERLYHECPKP SSMRRQQLIR ECPILSNIEP 60 KQIKVWFQNR RCREKQRKES FRLQAVNRKL TAMNKLLMEE NDRLQKQVSQ LVYENSYFRQ 120 QTQNAAITTT DTSCDSVVTS SRHHLTPQHP PRDASPAGLL SIAEETLAEF LSKATGTAVE 180 WVQMPGMKPG PDSIGIVAIS HGCAGVAARA CGLVGIEPAK VVEFLKDQPS WSRDCRSVDV 240 VSVLTTGNNG TVELLYMQLY APTTLAPARD LWLMRYTSVL EDGSLVVCER SLNNTQGGPT 300 MPSVQPFVRA EMLPSGYLIR PCDDGGSVIH IVYHMDLEPW SIPEVIRPLY ESSTMLAQRT 360 TMTALRHLRQ IAHEGSHSIA TGWGRRPAAL RALSQRLGRG FSEAINGFAD EGWSAASNDG 420 ADDVTILVKN YGFTNEFSMC NAVLCAKASM LLQNVSPAML LRFLREHRSE WGDSNMDAFS 480 AAALRANSCN FSVPHIGPFG GQVILPLAHS LDNEELLEVL KLENIGYYRD ALVPRELLLL 540 QLCTGLDENA VGTWSELIFA PIDASLADDA PLLPSGFRII PIDSIMDTCS PSNRTLDLAS 600 TLEVGPTGTR VCGDPSGRYS ATRSVMTIAF QFAFESHLQE NVASMARHYV RSIISSVQRV 660 ALALSPSRIA SLGGLCAPPG SPEVLMLARW ICESYRGHSG VELLKPTTEG NEPLLKSLWH 720 HSDAVVCCSV KDAPIFMFAN QAGLDMLETT SVALQEITPE QIFDDNGKKT LCNEFPQIIQ 780 KGFACVQGGL CVSSMGRPVS YEKAVAWKVL NDEDDTHCIC FMFVNWSFI* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
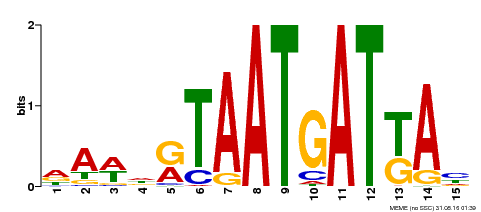
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020087984.1 | 0.0 | homeobox-leucine zipper protein ATHB-15-like | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A199VIA6 | 0.0 | A0A199VIA6_ANACO; Homeobox-leucine zipper protein ATHB-15 | ||||
| STRING | XP_008791633.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP374 | 38 | 197 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.2 | 0.0 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aco015166.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



