 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aco012822.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Bromeliaceae; Ananas
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 372aa MW: 40161.7 Da PI: 8.7722 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.4 | 1.3e-40 | 132 | 209 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+CqvegC++ l +ak+yh+rhkvC+vh+kap+v+v g eqrfCqqCsrfh +sefD++krsCrrrLa+hnerrrk+++
Aco012822.1 132 RCQVEGCDKSLVDAKDYHKRHKVCDVHAKAPKVIVLGAEQRFCQQCSRFHVISEFDDAKRSCRRRLAGHNERRRKSNT 209
6**************************************************************************975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 6.2E-33 | 129 | 194 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.257 | 130 | 207 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 8.11E-38 | 131 | 210 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 2.1E-31 | 133 | 206 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0010229 | Biological Process | inflorescence development | ||||
| GO:0010321 | Biological Process | regulation of vegetative phase change | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 372 aa Download sequence Send to blast |
MESGGGGGKS GHGGSSWGGS WELGNWTSAA VAANFPACYN AQGLEIAASA SASASRSAFP 60 DRAGVPLEGG DHAAEISQCA HRSFRSHHLT CLKLGKRHYY EEERLCGGAS AAAGKREKAA 120 AAAAAAGTPF PRCQVEGCDK SLVDAKDYHK RHKVCDVHAK APKVIVLGAE QRFCQQCSRF 180 HVISEFDDAK RSCRRRLAGH NERRRKSNTH DSLIRNPSSH SIEGTIMGPR FSHASNSPAR 240 ALSLLSSKAA SLPWVSPSDL SSRSSAALRE LIEENRAALL SRQLLSNRSD LPNQGPTNHI 300 AFPHWPQQNQ VVSQFPRLAD GWDHLQGAGA HVTLDLMQVP NPSFGAIPRR SRAKDGEEDC 360 SDIWKSWEDA HV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-30 | 133 | 206 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' (By similarity). May be involved in panicle development. {ECO:0000250}. | |||||
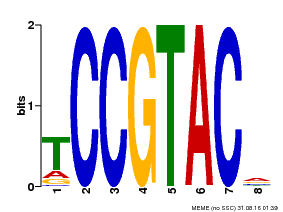
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00290 | DAP | Transfer from AT2G33810 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020084066.1 | 0.0 | squamosa promoter-binding-like protein 7 | ||||
| Swissprot | Q01JD1 | 1e-68 | SPL7_ORYSI; Squamosa promoter-binding-like protein 7 | ||||
| Swissprot | Q7XT42 | 9e-69 | SPL7_ORYSJ; Squamosa promoter-binding-like protein 7 | ||||
| TrEMBL | A0A199UT85 | 0.0 | A0A199UT85_ANACO; Squamosa promoter-binding-like protein 7 | ||||
| STRING | XP_008796104.1 | 1e-113 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP10303 | 34 | 39 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G33810.1 | 6e-36 | squamosa promoter binding protein-like 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aco012822.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



