 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aco004935.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Bromeliaceae; Ananas
|
||||||||
| Family | B3 | ||||||||
| Protein Properties | Length: 698aa MW: 78544.6 Da PI: 9.0446 | ||||||||
| Description | B3 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 54.1 | 2.7e-17 | 31 | 119 | 1 | 98 |
EEEE-..-HHHHTT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EEEEE- CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvvkvfr 98
ffkv+ ++++ +++pkkf+++++g+ s + l+++sg W++ + +k+++ ++l++GW eF a++++ +D + F+ +g+s+f v +++
Aco004935.1 31 FFKVM--VEDFTKC-MSMPKKFVDNFNGQ--ISNVVDLKTPSGDIWKIGV--TKSQDNIILQSGWTEFIAAHRIEKNDLLFFTCNGNSSF--EVAICD 119
78888..7788888.********998888..7889***************..*******************************9999999..888887 PP
| |||||||
| 2 | B3 | 46.2 | 8.5e-15 | 288 | 373 | 5 | 93 |
-..-H.HHHTT-EE--HHH.HTT---..--SEEEEEE.TTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..E CS
B3 5 ltpsd.vlksgrlvlpkkfaeehggkkeesktltled.esgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelv 93
+ ps+ + k++ l +pk fa++h +++s ++l+ + ++W+ +i+ + ++ + +GW++Fv++n+L+ gD+++F+ ++ +++ l+
Aco004935.1 288 MYPSNaAIKNCFLNIPKCFAKAHL--PRRSERIILKIpNKRETWTCSYIFATRLQG--IYGGWSDFVHDNNLRKGDTCLFEQMKGMKT-LT 373
555541468899***********5..3466678888834559********999999..569********************6654332.34 PP
| |||||||
| 3 | B3 | 62.1 | 9.1e-20 | 414 | 494 | 13 | 99 |
TT-EE--HHH.HTT---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..EEEEE-S CS
B3 13 sgrlvlpkkfaeehggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelvvkvfrk 99
++r+++pk+fa+++++k s + l+ +sg+ W+v + +k++++++l +GWkeFv+a++++e D ++Fkl+g+s+f v +f++
Aco004935.1 414 TQRMTIPKNFANRFSEK--ISEVVDLKPPSGNIWQVGV--TKSPDGIILVSGWKEFVEAHRVEESDLLLFKLIGTSSF--EVVIFDA 494
579********999755..7789***************..***********************************999..7777775 PP
| |||||||
| 4 | B3 | 48.9 | 1.2e-15 | 613 | 696 | 11 | 96 |
HHTT-EE--HHH.HTT---..--SEEEEEETTS.-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SS.SEE..EEEE CS
B3 11 lksgrlvlpkkfaeehggkkeesktltledesg.rsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgr.sefelvvkv 96
ks+ + +pkkfa++h ++ +s ++ l+ ++g ++W v +i +++ + +GWk+F+ +n+L+ gD+++F+l+++ +++v++
Aco004935.1 613 AKSNFVNIPKKFADAHLQP--RSHEIALKIPNGtNKWFVSYILGSDRRG--ICGGWKGFAADNNLRKGDICLFELMKNsGVPTMTVHI 696
4677899******999544..6779*****9999*******66555544..668**********************985666666665 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 3.0E-19 | 25 | 123 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.12E-19 | 28 | 126 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 12.605 | 28 | 121 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 5.55E-16 | 30 | 119 | No hit | No description |
| Pfam | PF02362 | 2.9E-14 | 31 | 119 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 4.9E-9 | 31 | 121 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 1.3E-20 | 280 | 377 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.28E-19 | 280 | 374 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.01E-15 | 282 | 377 | No hit | No description |
| SMART | SM01019 | 6.6E-9 | 284 | 380 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.435 | 285 | 380 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 1.7E-12 | 285 | 376 | IPR003340 | B3 DNA binding domain |
| SuperFamily | SSF101936 | 1.1E-21 | 398 | 496 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 14.677 | 402 | 495 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 1.1E-20 | 404 | 496 | IPR015300 | DNA-binding pseudobarrel domain |
| SMART | SM01019 | 8.7E-13 | 405 | 495 | IPR003340 | B3 DNA binding domain |
| CDD | cd10017 | 3.83E-17 | 413 | 493 | No hit | No description |
| Pfam | PF02362 | 2.0E-17 | 414 | 494 | IPR003340 | B3 DNA binding domain |
| Gene3D | G3DSA:2.40.330.10 | 6.5E-22 | 597 | 696 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.61E-21 | 597 | 696 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 6.32E-17 | 600 | 695 | No hit | No description |
| SMART | SM01019 | 1.6E-8 | 602 | 692 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.239 | 603 | 697 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 8.3E-14 | 614 | 696 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005773 | Cellular Component | vacuole | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 698 aa Download sequence Send to blast |
MKKISDKSCK ECEKWEEHFY WNHLDEHKMH FFKVMVEDFT KCMSMPKKFV DNFNGQISNV 60 VDLKTPSGDI WKIGVTKSQD NIILQSGWTE FIAAHRIEKN DLLFFTCNGN SSFEVAICDD 120 SGCEKASPFF AKKDARKTGE GDGATPAQTR NEGNRRRKVI ELCSSGSSSE NDVAPKISRK 180 TLCANSKTAN RIDCDVSSHG GAISGTNEKH KVPSKRPRGR PRKYIESSKG VEHVVKSESQ 240 ESCKCKEETP GSAPYYSSRS LQLTPSQDKK VSKLALALLP NPSFASVMYP SNAAIKNCFL 300 NIPKCFAKAH LPRRSERIIL KIPNKRETWT CSYIFATRLQ GIYGGWSDFV HDNNLRKGDT 360 CLFEQMKGMK TLTLTMFEGF ESCRECREWK EHFYWNHLDK QRTRFYTRMV GDFTQRMTIP 420 KNFANRFSEK ISEVVDLKPP SGNIWQVGVT KSPDGIILVS GWKEFVEAHR VEESDLLLFK 480 LIGTSSFEVV IFDASGHEKA SSLFATKKDP QMGESNAVSV EIISVENIHE VIAISSSTNS 540 SEVHAKPANC GVRRTGLDCG GRPNEREHEL HLELPYRSPP LPDWQEKKLS ALALAARPNP 600 TFVAIMHWSN ITAKSNFVNI PKKFADAHLQ PRSHEIALKI PNGTNKWFVS YILGSDRRGI 660 CGGWKGFAAD NNLRKGDICL FELMKNSGVP TMTVHII* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 208 | 217 | KHKVPSKRPR |
| 2 | 214 | 222 | KRPRGRPRK |
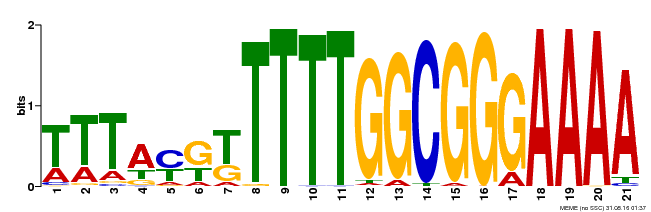
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00468 | DAP | Transfer from AT4G33280 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020092052.1 | 0.0 | B3 domain-containing protein Os03g0620400 | ||||
| TrEMBL | A0A199VWZ9 | 0.0 | A0A199VWZ9_ANACO; B3 domain-containing protein (Fragment) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP562 | 34 | 179 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G33280.1 | 2e-34 | B3 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aco004935.1 |



