 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Aco001898.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Bromeliaceae; Ananas
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 346aa MW: 38443.2 Da PI: 6.5389 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 56 | 1e-17 | 63 | 111 | 3 | 55 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+ GVr+++ +gr++AeIrdp++ + r++lg+f+ta+eAa a+++a+++++g
Aco001898.1 63 FLGVRRRP-WGRYAAEIRDPTT---KERHWLGTFDTAQEAALAYDRAALSMKG 111
56******.**********965...5************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00380 | 1.8E-32 | 62 | 125 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.666 | 62 | 119 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 3.24E-28 | 62 | 118 | No hit | No description |
| SuperFamily | SSF54171 | 4.32E-20 | 63 | 119 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 7.0E-28 | 63 | 119 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.3E-7 | 63 | 74 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 4.0E-11 | 65 | 111 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.3E-7 | 85 | 101 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010102 | Biological Process | lateral root morphogenesis | ||||
| GO:0010432 | Biological Process | bract development | ||||
| GO:0010451 | Biological Process | floral meristem growth | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 346 aa Download sequence Send to blast |
MSTSKTSNNT SMKTQVLQPT QHSSPSPPPP LSSSPSSSST SPTLSTSPSE RRGRRKQQEP 60 GRFLGVRRRP WGRYAAEIRD PTTKERHWLG TFDTAQEAAL AYDRAALSMK GTQARTNFMY 120 TSIDHHHLHH HNPFPSLHIP FLSPHLLHNK TFTIAPPPPP PPPQPCHHHN HQLIFSPQPM 180 QPTSESTPPH NSPSPVPDDQ LSSSLETEVD YFMFSDDSRS HGYLSSILPE SIFKSSDDDH 240 HQKHLPTSSA EEKTCSSYDH LEFNTQNTLS DHNSGFWGDH EPLWELSSIP HGLSKTSIGD 300 HFSMGEINGS LSFPPQGLDQ AYSPSDCVIS DAFDFGYSLF ENSLP* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 8e-20 | 53 | 118 | 5 | 71 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). Involved in the control of cell division patterns during the early lateral root primordium development. Acts downstream of auxin signaling (PubMed:17630277). Regulated by ARF7 and ARF19 in response to auxin. Co-acts with LBD16 and LBD18 to control lateral root development (PubMed:23749813). Involved in the determination of floral meristem identity and suppression of bract growth. Required for proper conversion of secondary inflorescences to flowers. Acts together with NPR5/BOP2 and NPR6/BOP1 to promote expression of LFY and AP1, two central regulators of floral meristem identity (PubMed:19482972). {ECO:0000250|UniProtKB:Q9LND1, ECO:0000269|PubMed:17630277, ECO:0000269|PubMed:19482972, ECO:0000269|PubMed:23749813}. | |||||
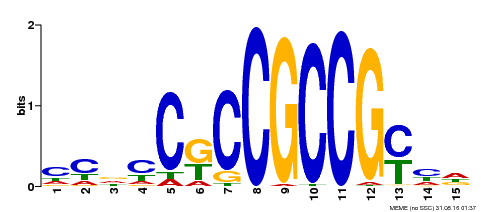
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00520 | DAP | Transfer from AT5G18560 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. {ECO:0000269|PubMed:17630277, ECO:0000269|PubMed:23749813}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020107857.1 | 0.0 | ethylene-responsive transcription factor ERF086-like | ||||
| Swissprot | Q6J9Q2 | 3e-44 | ERF86_ARATH; Ethylene-responsive transcription factor ERF086 | ||||
| TrEMBL | A0A199W217 | 0.0 | A0A199W217_ANACO; Ethylene-responsive transcription factor ERF086 | ||||
| STRING | XP_008802124.1 | 4e-57 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP10525 | 28 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G18560.1 | 2e-44 | ERF family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Aco001898.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



