 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Achn359611 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; Ericales; Actinidiaceae; Actinidia
|
||||||||
| Family | LBD | ||||||||
| Protein Properties | Length: 720aa MW: 80761 Da PI: 9.3223 | ||||||||
| Description | LBD family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF260 | 122.1 | 2.9e-38 | 523 | 624 | 1 | 101 |
DUF260 1 aCaaCkvlrrkCakdCvlapyfpaeq.pkkfanvhklFGasnvlkllkalpeeeredamsslvyeAearardPvyGavgvilklqqqleqlkaelallk 98
+C aCk+lrrkCa+dCv+apyf++eq p++fa++hk+FGasnv+kll ++p +r +a+ +++yeA+ar+rdPvyG+v++i++lqqq+++l+a+l ++k
Achn359611 523 PCGACKFLRRKCAADCVFAPYFCSEQgPAQFAAIHKVFGASNVSKLLLHVPVPDRCEAVVTIAYEAQARMRDPVYGCVAHIFALQQQVACLQAQLMQVK 621
7*************************99*******************************************************************9999 PP
DUF260 99 eel 101
++l
Achn359611 622 AQL 624
875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50064 | 12.852 | 1 | 58 | IPR001510 | Zinc finger, PARP-type |
| SuperFamily | SSF57716 | 1.57E-6 | 1 | 62 | No hit | No description |
| Gene3D | G3DSA:3.30.1740.10 | 5.9E-8 | 1 | 63 | IPR001510 | Zinc finger, PARP-type |
| SuperFamily | SSF57716 | 2.85E-18 | 66 | 149 | No hit | No description |
| Gene3D | G3DSA:3.30.1740.10 | 4.5E-21 | 66 | 153 | IPR001510 | Zinc finger, PARP-type |
| PROSITE profile | PS50064 | 16.001 | 70 | 142 | IPR001510 | Zinc finger, PARP-type |
| SMART | SM01336 | 4.8E-20 | 73 | 147 | IPR001510 | Zinc finger, PARP-type |
| Pfam | PF00645 | 1.0E-15 | 73 | 143 | IPR001510 | Zinc finger, PARP-type |
| SuperFamily | SSF142921 | 2.75E-6 | 277 | 324 | IPR008893 | WGR domain |
| Gene3D | G3DSA:1.20.142.10 | 7.4E-24 | 323 | 454 | IPR004102 | Poly(ADP-ribose) polymerase, regulatory domain |
| PROSITE profile | PS51060 | 18.536 | 344 | 472 | IPR004102 | Poly(ADP-ribose) polymerase, regulatory domain |
| SuperFamily | SSF47587 | 3.66E-19 | 345 | 454 | IPR004102 | Poly(ADP-ribose) polymerase, regulatory domain |
| Pfam | PF02877 | 8.3E-22 | 345 | 454 | IPR004102 | Poly(ADP-ribose) polymerase, regulatory domain |
| Pfam | PF00644 | 5.0E-10 | 452 | 516 | IPR012317 | Poly(ADP-ribose) polymerase, catalytic domain |
| SuperFamily | SSF56399 | 1.34E-9 | 452 | 516 | No hit | No description |
| Gene3D | G3DSA:3.90.228.10 | 1.7E-12 | 455 | 516 | IPR012317 | Poly(ADP-ribose) polymerase, catalytic domain |
| PROSITE profile | PS50891 | 22.649 | 522 | 624 | IPR004883 | Lateral organ boundaries, LOB |
| Pfam | PF03195 | 9.7E-37 | 523 | 621 | IPR004883 | Lateral organ boundaries, LOB |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006471 | Biological Process | protein ADP-ribosylation | ||||
| GO:0010311 | Biological Process | lateral root formation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003950 | Molecular Function | NAD+ ADP-ribosyltransferase activity | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 720 aa Download sequence Send to blast |
MWNHADCTLK KGNQIKAHVL PFYDHNLDWQ RSLDDVEGIE LLRWEDQEKI RKYVESGGPQ 60 NTPSAAVMEY GMEVSQTSRA TCRRCNQMIM KGEVRISTKP DGRGGKGLAW HHANCFMELF 120 PNTQVEKVSG WNNLSASDQA AVLALTQKLS NMRTKNLYTN WHRKWFKSQK AKKPSRTLPQ 180 PSNMPPGNQA ANEQHQSSNS KKLGDLKVSI IGLPKESMSI IKLSLEHRGF AIVSLSVISM 240 KRREGDMQDE WKSQIEGAGG QFHTKIKKDT NCLVVSGVLN DKNAEMKAAS YYILQIIHED 300 KGSCYVFRKW SQVGNEKIGG IKLAEMDYGV NKEVSKKKNL NGAGSLLAPP LKELIKMLFN 360 VEAYRICFRY IKFCRSAIME FEINMSEMPL GKLSKSSIQR GFEALTEIQN LVTSNVHDPS 420 YKESLIIDAS NRFFTVIPSI HPHVIRDEDD LKSKYMDKPP KGKHSTKGLG MKIPQKSEHA 480 KWRDEVVVPC GKPVASNIKA SELMFNEYIV YNTTQMAAGA GSPCGACKFL RRKCAADCVF 540 APYFCSEQGP AQFAAIHKVF GASNVSKLLL HVPVPDRCEA VVTIAYEAQA RMRDPVYGCV 600 AHIFALQQQV ACLQAQLMQV KAQLAQNLVD TRQVENPIWP GNMVGMPNFP AYANPTYMSL 660 SSISPQSSLE SIDRGNVDGM GVQERERREE FCFQICTKKR PSQTDLGELQ ALALRMMRN* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5ly0_A | 2e-34 | 519 | 624 | 7 | 111 | LOB family transfactor Ramosa2.1 |
| 5ly0_B | 2e-34 | 519 | 624 | 7 | 111 | LOB family transfactor Ramosa2.1 |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00309 | DAP | Transfer from AT2G42430 | Download |
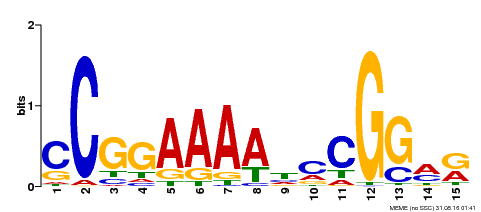
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| TrEMBL | A0A371HJR6 | 1e-122 | A0A371HJR6_MUCPR; Poly [ADP-ribose] polymerase (Fragment) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA43 | 24 | 669 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G42430.1 | 2e-50 | lateral organ boundaries-domain 16 | ||||



