 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Achn348581 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; Ericales; Actinidiaceae; Actinidia
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 611aa MW: 67473.6 Da PI: 6.6606 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 30.4 | 1e-09 | 217 | 273 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pseng...krkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++gr++A+++d + rk + g ++ +++Aa+ ++ a++k++g
Achn348581 217 SIYRGVTRHRWTGRYEAHLWDnSCR-RegqARKGRQ-GGYDKEDKAARSYDLAALKYWG 273
57*******************4444.2344336655.7799999*************98 PP
| |||||||
| 2 | AP2 | 49.5 | 1.1e-15 | 316 | 367 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s y+GV++++++grW A+I + +k +lg+f t+eeAa+a++ a+ k++g
Achn348581 316 SIYRGVTRHHQQGRWQARIGRVAG---NKDLYLGTFATEEEAAEAYDIAAIKFRG 367
57****************988532...5************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.24E-13 | 217 | 283 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.5E-7 | 217 | 273 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.95E-18 | 217 | 283 | No hit | No description |
| PROSITE profile | PS51032 | 17.042 | 218 | 281 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 3.4E-11 | 218 | 282 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.4E-20 | 218 | 287 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.7E-6 | 219 | 230 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 8.85E-12 | 316 | 375 | No hit | No description |
| Pfam | PF00847 | 5.6E-10 | 316 | 367 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.18E-17 | 316 | 376 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 4.8E-18 | 317 | 375 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.151 | 317 | 375 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.6E-31 | 317 | 381 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 4.7E-6 | 357 | 377 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010080 | Biological Process | regulation of floral meristem growth | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0035265 | Biological Process | organ growth | ||||
| GO:0048364 | Biological Process | root development | ||||
| GO:0060772 | Biological Process | leaf phyllotactic patterning | ||||
| GO:0060774 | Biological Process | auxin mediated signaling pathway involved in phyllotactic patterning | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 611 aa Download sequence Send to blast |
MAPESSNWLS FSLSPMEMLN SSSQAQMLQS TMKNVPFDGA SGDSQQYYFL DNFYGNGWEN 60 KAQEVTAQSI ADSTIFTTFL DSQIQHPPPK LEDFLGNDSS LVRYSDSRTE TQDSSLTHLY 120 DHSGSVYFAD HHDLKTVPAT AGFQAFSGNS GSEVDDSASV ARTQLVSTES GTELGFSQCP 180 NGALSLGVSA RGSEKALVSV DSESCKKISD TFGQRTSIYR GVTRHRWTGR YEAHLWDNSC 240 RREGQARKGR QGGYDKEDKA ARSYDLAALK YWGPTATTNF PVSSYTTEIE EMKHVTKQEF 300 IASLRRKSSG FSRGASIYRG VTRHHQQGRW QARIGRVAGN KDLYLGTFAT EEEAAEAYDI 360 AAIKFRGVNA VTNFEMNRYD VEAIMKSSLP IGGAAKRLKL SLEAEQKPSL NHDQQPPQCT 420 TSSSINFSTI IHPLAAVPCG IPFDTTARYH HNLFHLNTDG ASDPPGSAAA TPQSKAAIGV 480 GSRYGEKDMD NMSQKFRNEE LPETWCSQFF EKYKATKPYR LLLGSNLGDL AELVKDSTRN 540 SGQQTPQQLQ VMQSQIQSQQ QAAIRQPQQE HRNMSTVRAQ PVKVEDFQEL GGDASLKHDS 600 EENKLTSPPK * |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00504 | DAP | Transfer from AT5G10510 | Download |
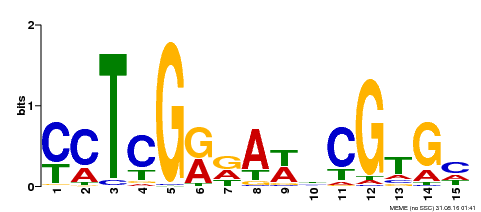
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028123293.1 | 0.0 | AP2-like ethylene-responsive transcription factor AIL6 | ||||
| Swissprot | Q52QU2 | 0.0 | AIL6_ARATH; AP2-like ethylene-responsive transcription factor AIL6 | ||||
| TrEMBL | A0A2R6S303 | 0.0 | A0A2R6S303_ACTCH; AP2-like ethylene-responsive transcription factor | ||||
| STRING | EOX91925 | 0.0 | (Theobroma cacao) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2580 | 24 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G10510.2 | 0.0 | AINTEGUMENTA-like 6 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



