 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Achn176661 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; Ericales; Actinidiaceae; Actinidia
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 821aa MW: 94598.6 Da PI: 6.9604 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 83.4 | 3.7e-26 | 89 | 192 | 1 | 91 |
FAR1 1 kfYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkk.............tekerrtraetrtgCkaklkvkkekdgkwevtkleleHn 86
+fY+eYA+++GF++ +++s++sk+++e+++++f Cs++g+++e +k+ t+++ +r+ +t+Cka+++vk++ dgkw ++++e+eHn
Achn176661 89 SFYQEYARSMGFNTAIQNSRRSKTSREFIDAKFACSRYGTKREYEKSlnqqrkrqgnkqeTDNSTGRRSCAKTDCKACMHVKRRPDGKWIIHRFEKEHN 187
5******************************************999988888887766554555559999***************************** PP
FAR1 87 Helap 91
Hel p
Achn176661 188 HELLP 192
**975 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.4E-23 | 89 | 192 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 4.3E-30 | 290 | 382 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.087 | 570 | 606 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 1.6E-4 | 577 | 604 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 5.8E-8 | 581 | 608 | IPR006564 | Zinc finger, PMZ-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009585 | Biological Process | red, far-red light phototransduction | ||||
| GO:0010218 | Biological Process | response to far red light | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 821 aa Download sequence Send to blast |
MDIDLRLPSG EHEKEDEEPS GIDNMLDGED KLQNGDEGNG NLVDVGDELN LADGRNMNSS 60 AVDMVVFKED TNLEPLAGME FESHGEAYSF YQEYARSMGF NTAIQNSRRS KTSREFIDAK 120 FACSRYGTKR EYEKSLNQQR KRQGNKQETD NSTGRRSCAK TDCKACMHVK RRPDGKWIIH 180 RFEKEHNHEL LPAQAVSEQT RKMYAAMARQ FAEYKSVVGL KNDSKNPFDK GRYLTLEAGE 240 AKILLEFFTQ MQIANSNFFY AIDISEDQRL KNLFWLDAKS RHDYINFCDV VSFDTTYVRN 300 KYKMSLALFI GVNQHYQFML LGCALVADES ASTFSWVLQT WLRAMGGQAP KVIITDQDNA 360 MKSVISEVFP STYHCFSLWH ILGKVSDTFS HLVKQHENFT EKLEKCIYRS WTHEEFEKRW 420 RKWVDKFELK ENEWIQSLLE DRTHWAPTYM KDVFLAGMYM PQRSESVNSF FDKYMHKKTS 480 VQEFLKQYES ILQDRYEEEA KADSDTWNKQ PALKSPSPFE KHVSGLYTHA VFKKFQTEVL 540 GAVACHPKRE RQDETTITFR VQDFEKNQEF IVTWNGLKSE VSCICRLFEF KGFLCRHALI 600 VLQICGLSYV PSQYILKRWT KDAKSRCLTG EASEQAQSRV QRYNDLCQRA MKLGEEGSLS 660 QDSYSFAFRA LEDAFGNCVS VNNSNKNLVE TGTSVTAGLL CIEEDNQSRS MSKTSKKKNP 720 TKKRKMNLEQ DVMTVGPQDS LQQMDKLSSR PVLDAYFGPQ QSMQGMVQLN LMAPTRDNYY 780 GTQQTIQGLG QLNSIAPNHD GYYSTQPSLH GLVYFARNVE * |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CACGCGC-3'. Activates the expression of FHY1 and FHL involved in light responses. When associated with PHYA, protects it from being recognized and degraded by the COP1/SPA complex. Positive regulator of chlorophyll biosynthesis via the activation of HEMB1 gene expression. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:12753585, ECO:0000269|PubMed:17012604, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:18715961, ECO:0000269|PubMed:22634759}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00078 | ChIP-seq | Transfer from AT3G22170 | Download |
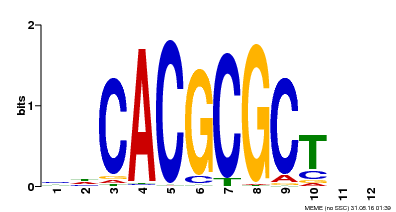
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated after exposure to far-red light. Subject to a negative feedback regulation by PHYA signaling. Up-regulated by white light. {ECO:0000269|PubMed:11889039, ECO:0000269|PubMed:18033885, ECO:0000269|PubMed:22634759}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_028080100.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Refseq | XP_028080101.1 | 0.0 | protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| Swissprot | Q9LIE5 | 0.0 | FHY3_ARATH; Protein FAR-RED ELONGATED HYPOCOTYL 3 | ||||
| TrEMBL | A0A2R6PUS0 | 0.0 | A0A2R6PUS0_ACTCH; Protein FAR-RED ELONGATED HYPOCOTYL like | ||||
| STRING | VIT_03s0017g01780.t01 | 0.0 | (Vitis vinifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA496 | 21 | 103 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G22170.2 | 0.0 | far-red elongated hypocotyls 3 | ||||



