 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AT3G13040.2 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 449aa MW: 50112.1 Da PI: 6.6479 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 99.7 | 1.9e-31 | 241 | 295 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k r+rWtpeLHe+Fv+av +L G+ekAtPk++++lm+v+gLt++hvkSHLQkYRl
AT3G13040.2 241 KSRMRWTPELHESFVKAVIKLEGPEKATPKAVKKLMNVEGLTIYHVKSHLQKYRL 295
68****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 9.921 | 238 | 298 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-29 | 239 | 296 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 3.23E-16 | 240 | 295 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.0E-23 | 241 | 296 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 5.8E-9 | 243 | 294 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 3.5E-22 | 331 | 378 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0000013 | anatomy | cauline leaf | ||||
| PO:0000037 | anatomy | shoot apex | ||||
| PO:0000230 | anatomy | inflorescence meristem | ||||
| PO:0000293 | anatomy | guard cell | ||||
| PO:0008019 | anatomy | leaf lamina base | ||||
| PO:0009005 | anatomy | root | ||||
| PO:0009006 | anatomy | shoot system | ||||
| PO:0009009 | anatomy | plant embryo | ||||
| PO:0009010 | anatomy | seed | ||||
| PO:0009025 | anatomy | vascular leaf | ||||
| PO:0009029 | anatomy | stamen | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0009031 | anatomy | sepal | ||||
| PO:0009032 | anatomy | petal | ||||
| PO:0009046 | anatomy | flower | ||||
| PO:0009047 | anatomy | stem | ||||
| PO:0009052 | anatomy | flower pedicel | ||||
| PO:0020030 | anatomy | cotyledon | ||||
| PO:0020038 | anatomy | petiole | ||||
| PO:0020100 | anatomy | hypocotyl | ||||
| PO:0020137 | anatomy | leaf apex | ||||
| PO:0025022 | anatomy | collective leaf structure | ||||
| PO:0025281 | anatomy | pollen | ||||
| PO:0001054 | developmental stage | vascular leaf senescent stage | ||||
| PO:0001078 | developmental stage | plant embryo cotyledonary stage | ||||
| PO:0001081 | developmental stage | mature plant embryo stage | ||||
| PO:0001185 | developmental stage | plant embryo globular stage | ||||
| PO:0004507 | developmental stage | plant embryo bilateral stage | ||||
| PO:0007064 | developmental stage | LP.12 twelve leaves visible stage | ||||
| PO:0007095 | developmental stage | LP.08 eight leaves visible stage | ||||
| PO:0007098 | developmental stage | LP.02 two leaves visible stage | ||||
| PO:0007103 | developmental stage | LP.10 ten leaves visible stage | ||||
| PO:0007115 | developmental stage | LP.04 four leaves visible stage | ||||
| PO:0007123 | developmental stage | LP.06 six leaves visible stage | ||||
| PO:0007611 | developmental stage | petal differentiation and expansion stage | ||||
| PO:0007616 | developmental stage | flowering stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 449 aa Download sequence Send to blast |
MYIKAIMNRH RLLSAATDEC NKKLGQACSS SLSPVHNFLN VQPEHRKTPF IRSQSPDSPG 60 QLWPKNSSQS TFSRSSTFCT NLYLSSSSTS ETQKHLGNSL PFLPDPSSYT HSASGVESAR 120 SPSIFTEDLG NQCDGGNSGS LLKDFLNLSG DACSDGDFHD FGCSNDSYCL SDQMELQFLS 180 DELELAITDR AETPRLDEIY ETPLASNPVT RLSPSQSCVP GAMSVDVVSS HPSPGSAANQ 240 KSRMRWTPEL HESFVKAVIK LEGPEKATPK AVKKLMNVEG LTIYHVKSHL QKYRLAKYMP 300 EKKEEKRTDN SEEKKLALSK SEADEKKKGA IQLTEALRMQ MEVQKQLHEQ LEVQRVLQLR 360 IEEHAKYLEK MLEEQRKTGR WISSSSQTVL SPSDDSIPDS QNMSKTKASS PQPPLPAENK 420 ASETEDDKCE SPQKRRRLEN IAESEDPKR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-23 | 241 | 298 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-23 | 241 | 298 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-23 | 241 | 298 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-23 | 241 | 298 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-23 | 241 | 298 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-23 | 241 | 298 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-23 | 241 | 298 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-23 | 241 | 298 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 433 | 437 | KRRRL |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| At.11993 | 0.0 | bud| flower| root| silique | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 30682425 | 0.0 | ||||
| Genevisible | 257855_at | 0.0 | ||||
| Expression Atlas | AT3G13040 | - | ||||
| AtGenExpress | AT3G13040 | - | ||||
| ATTED-II | AT3G13040 | - | ||||
| Function -- GeneRIF ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
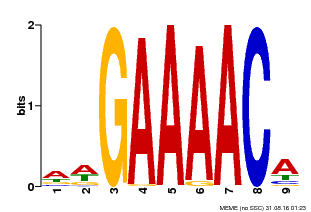
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | 27203113 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AT3G13040.2 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Interaction ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Intact With | |||||
| BioGRID | AT1G14920 | |||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| T-DNA Express | AT3G13040 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY050889 | 0.0 | AY050889.1 Arabidopsis thaliana unknown protein (At3g13040) mRNA, complete cds. | |||
| GenBank | AY096674 | 0.0 | AY096674.1 Arabidopsis thaliana unknown protein (At3g13040) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001319536.1 | 0.0 | myb-like HTH transcriptional regulator family protein | ||||
| Refseq | NP_566442.1 | 0.0 | myb-like HTH transcriptional regulator family protein | ||||
| Refseq | NP_974298.1 | 0.0 | myb-like HTH transcriptional regulator family protein | ||||
| Swissprot | Q949U2 | 0.0 | PHL6_ARATH; Myb family transcription factor PHL6 | ||||
| TrEMBL | A0A178VHF6 | 0.0 | A0A178VHF6_ARATH; Uncharacterized protein | ||||
| STRING | AT3G13040.1 | 0.0 | (Arabidopsis thaliana) | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | AT3G13040.2 |
| Entrez Gene | 820490 |
| iHOP | AT3G13040 |
| wikigenes | AT3G13040 |



