 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | AA55G00037 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Aethionemeae; Aethionema
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 676aa MW: 75317 Da PI: 5.8115 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 62.1 | 8.2e-20 | 47 | 101 | 2 | 56 |
T--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 2 rkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+k +++t++q++eLe++F ++++p++++r eL +klgL+ +q+k+WFqNrR+++k
AA55G00037 47 SKYHRHTSYQIQELESVFGECPHPNEKQRLELGRKLGLESTQIKFWFQNRRTQMK 101
67889***********************************************999 PP
| |||||||
| 2 | START | 134.9 | 8.9e-43 | 222 | 407 | 46 | 205 |
EEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEECTT...........................EEEEEEEEXXTTXX-SSX.EEEEEE CS
START 46 ealrasgvvdmvlallveellddkeqWdetla....kaetlevissg...........................galqlmvaelqalsplvp.Rdfvfv 112
e +r++g+v + +lve+l+++k qW e++a a+t+evis+g l +m+a++q++splvp R++ f+
AA55G00037 222 ECSRETGLVSISSLDLVETLMETK-QWAEMFAcivaVASTIEVISNGsngtrsgslhlvcssdtkdlifsqetdPLLGQMQAKFQVMSPLVPiREVKFL 319
679*********************.***********************************************9999*********************** PP
EEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 113 RyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqce 205
Ry++ql++ w++vdvS d++++ ++ +++lpSg++i++++ng skvtw+e+ +++++ +h l+r l++s + ga +w+atlqr c+
AA55G00037 320 RYCKQLKESFWAVVDVSYDVKKEDEK-----WCRRLPSGCIIQDMGNGCSKVTWIEQSEYDESYVHPLYRTLLSSAVGLGATRWLATLQRECQ 407
***********************996.....6679********************************************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.71E-19 | 31 | 103 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.9E-21 | 33 | 103 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 16.925 | 43 | 103 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 3.5E-17 | 44 | 107 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.01E-16 | 46 | 103 | No hit | No description |
| Pfam | PF00046 | 1.6E-17 | 47 | 101 | IPR001356 | Homeobox domain |
| SMART | SM00234 | 3.1E-23 | 177 | 408 | IPR002913 | START domain |
| CDD | cd08875 | 1.55E-83 | 196 | 407 | No hit | No description |
| SuperFamily | SSF55961 | 9.61E-25 | 196 | 269 | No hit | No description |
| Pfam | PF01852 | 2.6E-36 | 222 | 407 | IPR002913 | START domain |
| PROSITE profile | PS50848 | 30.827 | 225 | 411 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 9.61E-25 | 299 | 409 | No hit | No description |
| SuperFamily | SSF55961 | 2.88E-17 | 436 | 669 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 676 aa Download sequence Send to blast |
MKVLASPAKE KMKGNGDLNL KDDEFDSFDG AMSGDDKEEE APRKKKSKYH RHTSYQIQEL 60 ESVFGECPHP NEKQRLELGR KLGLESTQIK FWFQNRRTQM KTQLERHENV ILRQENEKLR 120 VENRYLKEAM RVPICSDCTG EVAPCEVSYD HQLKIENSKL KEELDRICTL TNRYIGSQNL 180 PVRPDFNGGS VMYESSMFME LAVKAMDELI KLADPGFVIV PECSRETGLV SISSLDLVET 240 LMETKQWAEM FACIVAVAST IEVISNGSNG TRSGSLHLVC SSDTKDLIFS QETDPLLGQM 300 QAKFQVMSPL VPIREVKFLR YCKQLKESFW AVVDVSYDVK KEDEKWCRRL PSGCIIQDMG 360 NGCSKVTWIE QSEYDESYVH PLYRTLLSSA VGLGATRWLA TLQRECQSLT TLFSCLNPVQ 420 DFAGLSPAGT KSALKLAKRM TFNFYSGITA SSSRRWEKLL AGNVGEDTRI LTRKNIDDPG 480 EPYGVILSAA TSLWLPVSHQ KLFDFLRDEK HRHHWDILCN GALMDDMLLV PKGERDGSCC 540 VSLLRAAGKD KTENSMLILQ ETWIDTSGAL VVYAPVDVSS MNGVMNGGDT ACVALLPSGF 600 SILPDGSSSS DQICSNGGLV HPESKENSSG GSLLTVGFQI LVNSLPTAIL NLDSLDTVNN 660 LISCTIHKIR SALHVP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
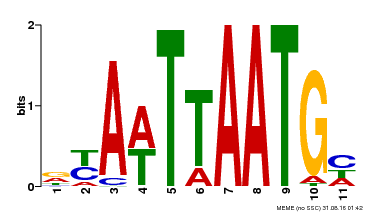
| UniProt | Probable transcription factor that binds to the DNA sequence 5'-GCATTAAATGC-3'. {ECO:0000269|PubMed:16778018}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00559 | DAP | Transfer from AT5G52170 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | AA55G00037 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_019091179.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein HDG7-like isoform X2 | ||||
| Swissprot | Q9LTK3 | 0.0 | HDG7_ARATH; Homeobox-leucine zipper protein HDG7 | ||||
| TrEMBL | A0A1P8BG00 | 0.0 | A0A1P8BG00_ARATH; Homeodomain GLABROUS 7 | ||||
| STRING | XP_010444530.1 | 0.0 | (Camelina sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1128 | 27 | 105 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52170.1 | 0.0 | homeodomain GLABROUS 7 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



