 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 678325266 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Utricularia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 812aa MW: 88701.5 Da PI: 6.1116 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 65.8 | 5.7e-21 | 138 | 193 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+k+l+L++rqVk+WFqNrR+++k
678325266 138 KKRYQRHTPQQIQELEALFKECPHPDEKQRLELSKRLSLETRQVKFWFQNRRTQMK 193
688999***********************************************999 PP
| |||||||
| 2 | START | 191 | 6e-60 | 336 | 554 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EEEEEEEECT CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....kaetleviss 87
ela++a++elvk+a+++ep+W +s e +n de++++f + + + +ea+r+sgvv+ ++ lve+l+d++ +W e+++ + +t++vis+
678325266 336 ELALSAMDELVKMAQSGEPLWIRSLetgtETLNHDEYTRNFNPCIGmrpngFITEASRESGVVIVNSSALVETLMDSN-KWAEMFPytvaRTTTTDVIST 434
57899************************************998889*******************************.********************* PP
T......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--.-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHH CS
START 88 g......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppesssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwl 180
g g lqlm aelq+lsplvp Rd f+R+++ql +g+w++vd+S+d ++ p +++lpSg+l+++++ng+skvtwveh+++++ +h l
678325266 435 GmggtrnGCLQLMHAELQVLSPLVPvRDIHFLRFCKQLAEGVWAVVDISIDAIRETPG------CRRLPSGCLVHDMPNGYSKVTWVEHTEYDETEVHVL 528
*******************************************************997......788********************************* PP
HHHHHHHHHHHHHHHHHHHTXXXXXX CS
START 181 lrslvksglaegaktwvatlqrqcek 206
+r+++ksgl +ga +wvatlqrqce+
678325266 529 YRPMIKSGLGFGAPRWVATLQRQCEC 554
************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.17E-20 | 121 | 195 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-21 | 124 | 189 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.831 | 135 | 195 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.0E-18 | 137 | 199 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 1.7E-18 | 138 | 193 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.93E-19 | 138 | 195 | No hit | No description |
| PROSITE pattern | PS00027 | 0 | 170 | 193 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 39.893 | 327 | 557 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 7.6E-35 | 329 | 554 | No hit | No description |
| CDD | cd08875 | 7.15E-119 | 331 | 553 | No hit | No description |
| Pfam | PF01852 | 8.7E-52 | 336 | 554 | IPR002913 | START domain |
| SMART | SM00234 | 2.2E-42 | 336 | 554 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 1.54E-24 | 575 | 799 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 812 aa Download sequence Send to blast |
MNFGDFLDSN NNILGASVGF GTDLPYANGN SHKIAIVAPP SSNGIINAMP SGVIAHSPHL 60 VSQNLSNKIF NSPGLSLALQ SGMDGQGEVA RTGENYDGSN VGGGKRSRDD DHESRSTSDN 120 FDGASGDDHD GTDKPSRKKR YQRHTPQQIQ ELEALFKECP HPDEKQRLEL SKRLSLETRQ 180 VKFWFQNRRT QMKTQLERHE NSLLRQENDK LRAENLAMRE AMRNPICTNC GGPAIMGEIS 240 LEEQQLRIEN ARLKDELDRV CVLAGKFLGR PVFSHNSSLK LGVGGNGYGD LSAHPSMLPL 300 GLPDFGSEMP IIPTTKQTMN AAAPIERSLE RSMYLELALS AMDELVKMAQ SGEPLWIRSL 360 ETGTETLNHD EYTRNFNPCI GMRPNGFITE ASRESGVVIV NSSALVETLM DSNKWAEMFP 420 YTVARTTTTD VISTGMGGTR NGCLQLMHAE LQVLSPLVPV RDIHFLRFCK QLAEGVWAVV 480 DISIDAIRET PGCRRLPSGC LVHDMPNGYS KVTWVEHTEY DETEVHVLYR PMIKSGLGFG 540 APRWVATLQR QCECLAILMS SAAQDKDPHS GAITPGGRRS MLKLAQRMTN NFCAGVCAST 600 VHNWDKLRTE NVDDDVRVMT RKSVDDPGEP QGIVLSAATS VWLPVTPQRL FDFLRNEHLR 660 SEWDILSNGG PMQEMAHIAK GQDHGNCVSL LRASAMNANQ SNMLILQETC VDPAGSLVVY 720 APVDIPAMHV VMNGGDSAYV ALLPSGFAIV PDGRGAGDGA GGGFQGSLLT VAFQILVNSL 780 PTAKLTVESV ETVNNLISCT VQKIKAALHC EG |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
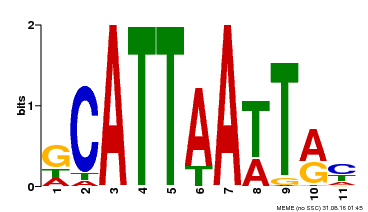
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 678325266 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011093167.1 | 0.0 | homeobox-leucine zipper protein ANTHOCYANINLESS 2 isoform X2 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A2G9HWV6 | 0.0 | A0A2G9HWV6_9LAMI; Transcription factor DLX | ||||
| STRING | XP_009602856.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1626 | 24 | 65 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G00730.1 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



