 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 678324670 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lentibulariaceae; Utricularia
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 725aa MW: 78600.4 Da PI: 6.4048 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 25.8 | 2.4e-08 | 24 | 50 | 1 | 28 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIa 28
r rWT+eE+ ++++a k++G W +I
678324670 24 RERWTEEEHNRFLEALKLHGRA-WQRIE 50
78******************88.*9995 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 5.77E-14 | 18 | 89 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.031 | 19 | 88 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 5.4E-10 | 22 | 86 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 7.9E-6 | 23 | 79 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.0E-6 | 23 | 86 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.1E-5 | 24 | 50 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 4.69E-7 | 26 | 84 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010243 | Biological Process | response to organonitrogen compound | ||||
| GO:0042754 | Biological Process | negative regulation of circadian rhythm | ||||
| GO:0043496 | Biological Process | regulation of protein homodimerization activity | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0048574 | Biological Process | long-day photoperiodism, flowering | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 725 aa Download sequence Send to blast |
MDPCFSGEDL VIKTRKPYTI TKQRERWTEE EHNRFLEALK LHGRAWQRIE EARLPFISIT 60 LDIGAEHIGT KTAIQIRSHA QKFFTKLEKE ATVKGTGNAL DIEIPPPRPK RKPANPYPRK 120 KCEVPALQLG LNDSKSSDTT STSLSMAFVK ESEDLSDDAK QQNSDHEKDD SITALNLDMV 180 APCTSPTSTN SSLESETPER SCTFRQFVPL VNGSHNQKGS AESLFSAKPT GSHAEQAQQK 240 RLLNDNILNS MVPMGLCPSP EESSHGEMVH DPIDETSHNC PRNVPVHILG GSCGSRQHPP 300 NMPCMKSVPN QTGFHGDQST FINAAASGTS ENGANGAAIP PQYLHCHPFF APMMNTEDYQ 360 SLQQMSSTFS GLVFSALAQN AALYAAASFA STMWPWLNLG GAPVEPSANS SGIPQPRPMN 420 PTSSMAAIAG ATVAAATAWW AANGMFPLGP QYHAGFACSP PSAAAAQTAG SQSGNVNGEA 480 RESGADPTLQ CQQPDGKLFN ALHSASKPPT SSSSDSEASE AEKLNGRLNP SGTAEAEKSS 540 GSNVSINEKQ MDRFSSGSNT LSSSEVEPDA TKMNEENHME TMEVTEQPNG VPPPTIDLSS 600 RRSRSNNSLN DPWKEVSQEG RLAFQALFSR EVLPQSFPPS HVLTNKLKKH CSGCREDGDG 660 ASIFKPDEEK GSSFNIGKKH EGPLNQELGH TKAKDYRTGF RLYKRCSMEA KEIPEEHKDS 720 KKLRL |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 107 | 120 | RPKRKPANPYPRKK |
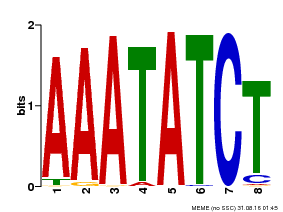
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00103 | PBM | Transfer from AT2G46830 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 678324670 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020553443.1 | 1e-160 | protein LHY-like | ||||
| TrEMBL | A0A2G9H753 | 1e-155 | A0A2G9H753_9LAMI; Uncharacterized protein | ||||
| STRING | Migut.N01518.1.p | 1e-137 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA9710 | 13 | 15 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46830.1 | 2e-37 | circadian clock associated 1 | ||||



