 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 676736652 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Sisymbrieae; Sisymbrium
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 508aa MW: 55666.7 Da PI: 9.5685 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 37.6 | 5.2e-12 | 69 | 113 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+ E++++++a +++ + Wk+I + +g +t q++s+ qky
676736652 69 RESWTEPEHDKFLEALQLFDRD-WKKIEAFIG-SKTVIQIRSHAQKY 113
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 1.12E-15 | 63 | 119 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 13.758 | 64 | 118 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.6E-6 | 67 | 119 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 4.5E-18 | 67 | 116 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 9.6E-10 | 68 | 116 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.2E-9 | 69 | 113 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.12E-7 | 71 | 114 | No hit | No description |
| Pfam | PF03501 | 1.9E-42 | 329 | 421 | IPR005326 | Plectin/S10, N-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 508 aa Download sequence Send to blast |
MVSRNPNPAE GYFLDPNGMT VPGIGPFTAA VSSSPTTSST AVGVADATVS SSEDLSKKIR 60 KPYTITKSRE SWTEPEHDKF LEALQLFDRD WKKIEAFIGS KTVIQIRSHA QKYFLKVQKS 120 GTGEHLPPPR PKRKAAHPYP QKAHKNAQPQ VPASFKSTPE PNDSSYMFRP ESSSMLMTSP 180 TTPPAVAPWT NNVQTISFTP LPKGCCLFFN ILIEAAGAGA NNNCSSSSEN TPRPRANKDT 240 NEQGNLGQSL RVLPDFAQVY SFIGSVFDPH ASNHLQKLKK MDPIDVETVL LLMRNLSINL 300 SSPDFEDHRR LLSSYDIGSE TATDHGGIIS EENRREICKY LFKEGVCFAK KDFNLAKHPL 360 IESVPNLQVI KLMQSFKSKE YVRETFAWMH YYWFLTNEGI EFLRTYLNLP SDVVPATLKK 420 SAKPIGRPFG GPPGDRPRGP RFEGGDRPRY GDRDGYRGGP RAGGEFGGDK SGAPADYQPS 480 FQGSGGRPGF GRGAGGYGAA APSGSGLP |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4v7e_BK | 2e-70 | 328 | 484 | 2 | 159 | 40S ribosomal protein S10E |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
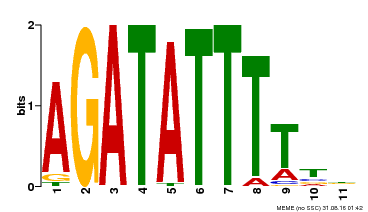
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00560 | DAP | Transfer from AT5G52660 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 676736652 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AY065217 | 1e-175 | AY065217.1 Arabidopsis thaliana unknown protein (At5g52650) mRNA, complete cds. | |||
| GenBank | AY122932 | 1e-175 | AY122932.1 Arabidopsis thaliana unknown protein (At5g52650) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_006401794.1 | 0.0 | protein REVEILLE 6 isoform X1 | ||||
| Refseq | XP_024012565.1 | 0.0 | protein REVEILLE 6 isoform X2 | ||||
| Swissprot | Q8H0W3 | 1e-177 | RVE6_ARATH; Protein REVEILLE 6 | ||||
| TrEMBL | A0A2P5FGQ1 | 0.0 | A0A2P5FGQ1_TREOI; GAMYB transcription factor | ||||
| STRING | Lus10038846 | 0.0 | (Linum usitatissimum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1290 | 28 | 93 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G52660.1 | 1e-148 | MYB_related family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



