 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 676715014 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Sisymbrieae; Sisymbrium
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 550aa MW: 63482.7 Da PI: 6.3279 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 35.9 | 1.6e-11 | 92 | 135 | 4 | 47 |
XCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 4 lkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkk 47
+k rr+++NReAAr+sR+RKka++++Lee +L++ ++L k
676715014 92 DKTKRRLAQNREAARKSRLRKKAYVQQLEESRLKLSQLEQELEK 135
7999**************************98888888776654 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 1.9E-5 | 84 | 147 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.335 | 91 | 135 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 2.1E-8 | 92 | 136 | No hit | No description |
| Pfam | PF00170 | 8.1E-8 | 92 | 135 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.37E-6 | 93 | 135 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 96 | 111 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 3.3E-28 | 180 | 254 | IPR025422 | Transcription factor TGA like domain |
| PROSITE profile | PS50076 | 15.397 | 354 | 427 | IPR001623 | DnaJ domain |
| SMART | SM00271 | 9.6E-10 | 355 | 419 | IPR001623 | DnaJ domain |
| PRINTS | PR00625 | 2.6E-12 | 358 | 376 | IPR001623 | DnaJ domain |
| CDD | cd06257 | 3.69E-11 | 362 | 416 | IPR001623 | DnaJ domain |
| Pfam | PF00226 | 1.6E-16 | 362 | 424 | IPR001623 | DnaJ domain |
| Gene3D | G3DSA:1.10.287.110 | 1.1E-17 | 363 | 426 | IPR001623 | DnaJ domain |
| SuperFamily | SSF46565 | 1.44E-17 | 363 | 425 | IPR001623 | DnaJ domain |
| PRINTS | PR00625 | 2.6E-12 | 376 | 391 | IPR001623 | DnaJ domain |
| PRINTS | PR00625 | 2.6E-12 | 399 | 419 | IPR001623 | DnaJ domain |
| PROSITE pattern | PS00636 | 0 | 404 | 423 | IPR018253 | DnaJ domain, conserved site |
| PRINTS | PR00625 | 2.6E-12 | 445 | 464 | IPR001623 | DnaJ domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 550 aa Download sequence Send to blast |
MMSSSPTQLA SLRDMGIYEP FQQIVSWGNV FKSDINDHSP NTASSSVIQV DARIDDHNSL 60 IKSNYASSSH NQMEAEPSSN DHQDDDDGRI PDKTKRRLAQ NREAARKSRL RKKAYVQQLE 120 ESRLKLSQLE QELEKAKQQG LCVRNSSDSS YLGPSGSMNT RIAAFEMEYS HWLEEQSRRV 180 SEIRRALQAH ISDIELRMLV ESSLNHYANL FRMKADAAKA DVFFLISGMW RTSTERFFQW 240 IGGFRPSELL NVVMPYLQPL TDQQILEVRN LQQSSQQAED ALSQGIDKLQ QSLVENIVVD 300 AVIESTDYPP HMAAALENLQ ALEGFVNQAD HLRQQTLQQM AKILTTRQAA RGLVLFWFLS 360 TELERDAEEE QIKVAYRRLA KYYHPDVYDG KGTLEEGETA EGRFIKIQAA YELLMDTEKR 420 RQYDMDNRVN PMKASQAWME WLIKKRKAFD QRGDMAIAAW AEQQQLEINL RARRLSRSKV 480 DPEEERKILE KEKKASRELV NSTLKRHTLV LKKRDLMRKK AEEDKKKLIT QLLAAEGLEL 540 DTEEEEEAPK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specifically to the DNA sequence 5'-TGACG-3'. Recognizes ocs elements like the as-1 motif of the cauliflower mosaic virus 35S promoter. Binding to the as-1-like cis elements mediate auxin- and salicylic acid-inducible transcription. May be involved in the induction of the systemic acquired resistance (SAR) via its interaction with NPR1 (By similarity). {ECO:0000250, ECO:0000269|PubMed:12953119}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00247 | DAP | Transfer from AT1G77920 | Download |
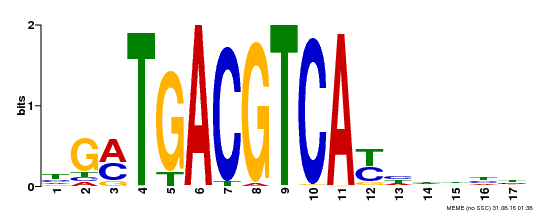
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 676715014 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK353384 | 0.0 | AK353384.1 Thellungiella halophila mRNA, complete cds, clone: RTFL01-35-N08. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010471992.1 | 0.0 | PREDICTED: transcription factor TGA7 | ||||
| Swissprot | Q93ZE2 | 0.0 | TGA7_ARATH; Transcription factor TGA7 | ||||
| TrEMBL | A0A3P6BCZ9 | 0.0 | A0A3P6BCZ9_BRACM; Uncharacterized protein | ||||
| STRING | Bo6g081910.1 | 0.0 | (Brassica oleracea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3061 | 27 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G77920.1 | 0.0 | bZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



