 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 60155 | ||||||||
| Common Name | MICPUCDRAFT_60155 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Mamiellaceae; Micromonas; Micromonas pusilla
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 509aa MW: 50762.6 Da PI: 9.779 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 46.8 | 7.3e-15 | 102 | 141 | 13 | 55 |
AP2 13 grWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
+W+AeIrdp + r +r +lg+f+taeeAa+a++aa++ ++g
60155 102 RSWAAEIRDP---N-RgARLWLGTFDTAEEAARAYDAAARHIRG 141
68*******8...3.35***********************9998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 2.42E-12 | 48 | 149 | No hit | No description |
| SMART | SM00380 | 5.0E-24 | 49 | 155 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.48 | 49 | 149 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.3E-10 | 50 | 61 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.57E-16 | 100 | 149 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 3.3E-24 | 100 | 149 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 2.6E-8 | 102 | 141 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 5.3E-10 | 115 | 131 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0034059 | Biological Process | response to anoxia | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071456 | Biological Process | cellular response to hypoxia | ||||
| GO:0071497 | Biological Process | cellular response to freezing | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 509 aa Download sequence Send to blast |
MSAAVAGNMH SPTDSLPPAN VEDDDLDDLD DDDDDPKAVT KTKKTTSSIY RGVRQRPWGR 60 EPLDDWHRGA RKLSPRKYRD VVEFSSVSDA LPRSLPRAVP SRSWAAEIRD PNRGARLWLG 120 TFDTAEEAAR AYDAAARHIR GPNARTNFQL APGEVPPPFV LPDPPTGGRG GRGRGDGPDS 180 PGGGRGGRGG RGGPHVAKKG RGAGAGMPKT GVGLGGGGYG VGGGGGGGFL GAFAGAHSNA 240 FKPSVVGASH GRFNATAHGG THTANDLSAL VNANLALAQQ NLSMAGAHGG HGAGAGSGSD 300 GGFIRLGGAG VGGGNHHHHH VVGEIPRIHN SKRERAPSNP FLGTSFGVAG SFGSVFGHDY 360 LSSTPDGGLG FPSHFGSSPG AGGELHGGSH HGRSGGGASH EGSHHGRSGG GGLGGGGFSL 420 GAAIKDDFEV GSLGAAGGLD DSVMPIGSME LGSPLDSFGD LMGSLPKGGS LGLKNSLGGG 480 NKNGAGGLPF GRSSQGDGRT ATDALKMR* |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00230 | DAP | Transfer from AT1G72360 | Download |
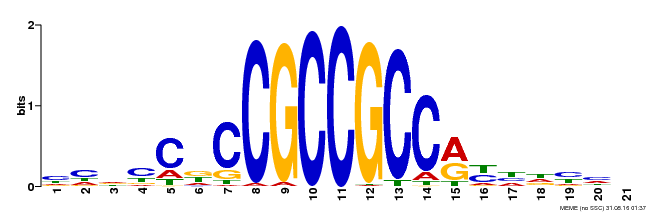
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_003060393.1 | 0.0 | erf-like protein | ||||
| TrEMBL | C1MXG9 | 0.0 | C1MXG9_MICPC; Erf-like protein | ||||
| STRING | XP_003060393.1 | 0.0 | (Micromonas pusilla) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP132 | 15 | 40 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G11140.1 | 9e-11 | cytokinin response factor 1 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 60155 |
| Entrez Gene | 9686129 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



