 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 486333 | ||||||||
| Common Name | ARALYDRAFT_486333 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 249aa MW: 28275.8 Da PI: 9.4261 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 97.6 | 5.1e-31 | 24 | 73 | 1 | 50 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeys 50
krien++nrqvtf+kRrng+lKKA+ELSvLCdaeva++ifs++g+lyey+
486333 24 KRIENTTNRQVTFCKRRNGLLKKAYELSVLCDAEVALVIFSTRGRLYEYA 73
79***********************************************8 PP
| |||||||
| 2 | K-box | 115.7 | 4.3e-38 | 95 | 188 | 7 | 100 |
K-box 7 ksleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkklee 100
s++ea+++++qqe++kL+++i+ +q+++Rh++Ge+L+sL+lkeL++Le +Lek+++++RskKnell+++ie++qk+e elq++n++Lr+k++e
486333 95 PSVTEANTQYYQQEASKLRRQIRDIQNSNRHIVGESLGSLNLKELKNLEGRLEKGISRVRSKKNELLVAEIEYMQKREMELQHNNMYLRAKIAE 188
45899**************************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00432 | 9.1E-40 | 16 | 75 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 33.389 | 16 | 76 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 1.04E-43 | 17 | 90 | No hit | No description |
| SuperFamily | SSF55455 | 1.96E-32 | 17 | 88 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.2E-32 | 18 | 38 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 18 | 72 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 3.3E-26 | 25 | 72 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.2E-32 | 38 | 53 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.2E-32 | 53 | 74 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 1.8E-27 | 100 | 186 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 15.15 | 102 | 192 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 249 aa Download sequence Send to blast |
MEEGGSSHDG ESSKKIGRGK IEIKRIENTT NRQVTFCKRR NGLLKKAYEL SVLCDAEVAL 60 VIFSTRGRLY EYANNSVRGT IERYKKACSD AVNPPSVTEA NTQYYQQEAS KLRRQIRDIQ 120 NSNRHIVGES LGSLNLKELK NLEGRLEKGI SRVRSKKNEL LVAEIEYMQK REMELQHNNM 180 YLRAKIAEGA RLNPEQQESS VIQGTTVYES GVSSHDQSQH HNRNYIPVNL LEPNQQFSGQ 240 DQPPLQLV* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1egw_A | 4e-20 | 17 | 84 | 1 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1egw_B | 4e-20 | 17 | 84 | 1 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1egw_C | 4e-20 | 17 | 84 | 1 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1egw_D | 4e-20 | 17 | 84 | 1 | 68 | MADS BOX TRANSCRIPTION ENHANCER FACTOR 2, POLYPEPTIDE A |
| 1tqe_P | 4e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_Q | 4e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_R | 4e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 1tqe_S | 4e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 5f28_A | 4e-20 | 16 | 84 | 1 | 69 | MEF2C |
| 5f28_B | 4e-20 | 16 | 84 | 1 | 69 | MEF2C |
| 5f28_C | 4e-20 | 16 | 84 | 1 | 69 | MEF2C |
| 5f28_D | 4e-20 | 16 | 84 | 1 | 69 | MEF2C |
| 6c9l_A | 4e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_B | 4e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_C | 4e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_D | 4e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_E | 4e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| 6c9l_F | 4e-20 | 16 | 84 | 1 | 69 | Myocyte-specific enhancer factor 2B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. Interacts genetically with TT16/AGL32 in a partially antagonistic manner during flower development. Is essential for the coordination of cell divisions in ovule, seed coat development and endosperm formation (PubMed:27776173). {ECO:0000269|PubMed:27776173}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00093 | SELEX | Transfer from AT3G58780 | Download |
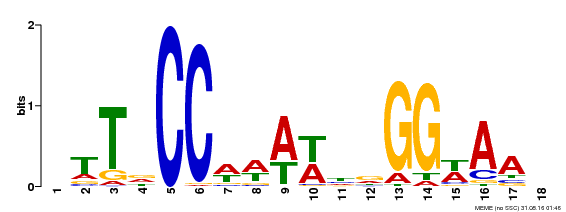
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 486333 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | ATHAGL1A | 0.0 | M55550.1 Arabidopsis thaliana transcription factor (AGL1), complete cds. | |||
| GenBank | AY086196 | 0.0 | AY086196.1 Arabidopsis thaliana clone 22339 mRNA, complete sequence. | |||
| GenBank | JQ973091 | 0.0 | JQ973091.1 Arabidopsis thaliana Shatterproof (SHP1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002876479.1 | 0.0 | agamous-like MADS-box protein AGL1 isoform X2 | ||||
| Swissprot | P29381 | 0.0 | AGL1_ARATH; Agamous-like MADS-box protein AGL1 | ||||
| TrEMBL | D7LW76 | 0.0 | D7LW76_ARALL; Uncharacterized protein | ||||
| STRING | fgenesh2_kg.5__2325__AT3G58780.1 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM3341 | 23 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G58780.1 | 1e-164 | MIKC_MADS family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 486333 |
| Entrez Gene | 9314308 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



