 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 462863802 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Eragrostideae; Eragrostidinae; Eragrostis
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 237aa MW: 25873.1 Da PI: 7.4523 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.4 | 8.4e-19 | 23 | 73 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.psengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s++ GVr+++ +grWvAeI+d + r +lg+f taeeAa+a+++a+ l+g
462863802 23 SKFVGVRQRP-SGRWVAEIKDtT-----HkIRMWLGTFETAEEAARAYDEAACLLRG 73
79********.7*********33.....35**********************99998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 1.37E-19 | 23 | 81 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 2.7E-13 | 23 | 73 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 1.18E-31 | 23 | 81 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 3.3E-28 | 24 | 81 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.823 | 24 | 81 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 1.2E-35 | 24 | 87 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.4E-8 | 25 | 36 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.4E-8 | 47 | 63 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000302 | Biological Process | response to reactive oxygen species | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0035865 | Biological Process | cellular response to potassium ion | ||||
| GO:0048528 | Biological Process | post-embryonic root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 237 aa Download sequence Send to blast |
MELQFQKPQQ SQQVAGTNKA ARSKFVGVRQ RPSGRWVAEI KDTTHKIRMW LGTFETAEEA 60 ARAYDEAACL LRGANTRTNF AGSSPPDSPL AIRALLNHKK LKKNHGGTVT ATSTTGPSSS 120 VTSSTTINFA MIDDDQVRTP NLPAARNLTE EAYLISGSED QFQLVASQPW ALNASLPPTD 180 ACSAMMTDQQ DKIKPKKESP ASPHAMDQAF YDTGNDPSDS LWDLPPICPL SCRSLMY |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5wx9_A | 5e-15 | 20 | 80 | 10 | 71 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in symbiotic nodule signaling in response to rhizobial Nod factors (NFs) (PubMed:17449807, PubMed:17827349). Binds to the GCC-box (NF-responsive box) of ENOD11 promoter. Acts as transcriptional activator of NF-responsive box-containing target gene promoters in root hairs (PubMed:17827349). Functions as a transcriptional regulator required for root infection by symbiotic rhizobia, infection thread (IT) formation and maintenance, and nodule development. Necessary for NF-induced gene expression and spontaneous nodulation activated by CCAMK. Functions downstream of CCAMK to activate nodulation gene expression (PubMed:17449807). Involved in early stages of root nodule development. Functions redundantly with ERN2. Is essential with ERN2 for the initiation of root hair infection, and nodule organogenesis and development. Required for accurate expression of the NF signaling genes ENOD11 and ENOD12 (PubMed:23077241, PubMed:27208242). {ECO:0000269|PubMed:17449807, ECO:0000269|PubMed:17827349, ECO:0000269|PubMed:23077241, ECO:0000269|PubMed:27208242}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00521 | DAP | Transfer from AT5G19790 | Download |
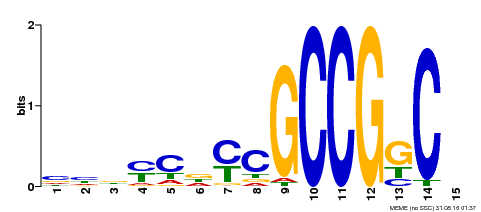
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 462863802 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced in roots from 1 to 21 day after inoculation with Sinorhizobium meliloti (PubMed:17449807). Induced by Nod factors in root hairs (PubMed:17827349). {ECO:0000269|PubMed:17449807, ECO:0000269|PubMed:17827349}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AP004273 | 7e-65 | AP004273.2 Oryza sativa Japonica Group genomic DNA, chromosome 7, PAC clone:P0431A02. | |||
| GenBank | AP014963 | 7e-65 | AP014963.1 Oryza sativa Japonica Group DNA, chromosome 7, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012615 | 7e-65 | CP012615.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 7 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002459520.2 | 4e-76 | ethylene-responsive transcription factor RAP2-11 isoform X1 | ||||
| Swissprot | A2Q5W1 | 5e-54 | ERN1_MEDTR; Ethylene-responsive transcription factor ERN1 | ||||
| TrEMBL | A0A1E5VKR9 | 3e-76 | A0A1E5VKR9_9POAL; Uncharacterized protein | ||||
| STRING | LPERR07G05890.1 | 2e-75 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3900 | 32 | 73 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G19790.1 | 4e-27 | related to AP2 11 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



