 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 462849714 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Eragrostideae; Eragrostidinae; Eragrostis
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 386aa MW: 40934.9 Da PI: 8.3909 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 69.2 | 6.4e-22 | 289 | 351 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
e+elkr+rrkq+NRe+ArrsR+RK+ae+eeL++++++L++eN++L++e+++++ke+++l s++
462849714 289 ERELKRQRRKQSNRESARRSRLRKQAECEELAQRAEVLKQENSSLRDEVNRIRKEYEELLSKN 351
89**********************************************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF07777 | 2.9E-29 | 1 | 98 | IPR012900 | G-box binding protein, multifunctional mosaic region |
| Pfam | PF16596 | 6.7E-38 | 137 | 264 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 8.5E-19 | 283 | 348 | No hit | No description |
| SMART | SM00338 | 3.1E-21 | 289 | 353 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 2.4E-20 | 289 | 351 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 13.518 | 291 | 354 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 5.38E-11 | 292 | 348 | No hit | No description |
| CDD | cd14702 | 6.16E-25 | 294 | 344 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 296 | 311 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 386 aa Download sequence Send to blast |
MGSSGTDTPT KASKPSATQE QQPPATSSTA TPAVYPDWSS FQGYPPIPPH GFFPSPVASS 60 PQGHPYMWGA QPMIPPYGTP PPPYVMYPPG VYAHPSMPPG AHPFAPYAMT SPNGNADASG 120 AAAAAGETDG KASEGKDKSP TKRSKGSLGS LNMLTGKSPT EHGKTSGASA NGAVSQSGES 180 GSESSSEGSE GNSQNDSHHK GSGHEQDGDV RSSQNGVSRS PSEGKLNQTM AIMPMPSTGP 240 VPGPTTNLNI GMDYWGANTA SSTPAVHAKG TLTTVPGAVV PGEQWIQDER ELKRQRRKQS 300 NRESARRSRL RKQAECEELA QRAEVLKQEN SSLRDEVNRI RKEYEELLSK NNSLKEKLGD 360 KQYKTDETGL DNKLQHSGDD VRKKGN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 305 | 311 | RRSRLRK |
| 2 | 305 | 312 | RRSRLRKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in responses to fungal pathogen infection and abiotic stresses. {ECO:0000269|Ref.1}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00291 | DAP | Transfer from AT2G35530 | Download |
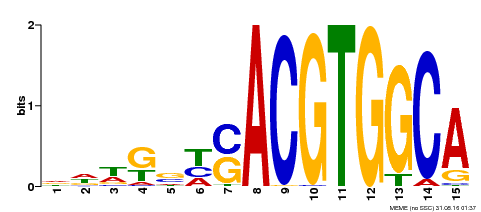
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 462849714 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced during incompatible interaction with the fungal pathogen Puccinia striiformis (Ref.1). Induced by abscisic acid (ABA), ethylene, cold stress, salt stress and wounding (Ref.1). {ECO:0000269|Ref.1}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | NP_001131383.1 | 0.0 | uncharacterized protein LOC100192709 | ||||
| Refseq | XP_008672076.1 | 0.0 | putative bZIP transcription factor superfamily protein isoform X1 | ||||
| Refseq | XP_008672077.1 | 0.0 | putative bZIP transcription factor superfamily protein isoform X1 | ||||
| Swissprot | B6E107 | 0.0 | BZP1B_WHEAT; bZIP transcription factor 1-B | ||||
| TrEMBL | B4FCD1 | 0.0 | B4FCD1_MAIZE; BZIP transcription factor 16 | ||||
| STRING | Pavir.Cb01814.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2147 | 37 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G35530.1 | 5e-82 | basic region/leucine zipper transcription factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



