 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 336483 | ||||||||
| Common Name | ARALYDRAFT_336483 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 548aa MW: 61621 Da PI: 4.6486 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 157.2 | 6.8e-49 | 17 | 143 | 2 | 128 |
NAM 2 ppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.k.kvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevlskkgelvgl 102
+pGfrFhPtdeelv++yLk+k+ +k+l++ +vi vd+yk++P++Lp + ++k+++++w++F++r++ky++ r++r t++gyWkatgkd+ + +++ vgl
336483 17 APGFRFHPTDEELVMYYLKRKICRKRLRV-NVIGVVDVYKMDPEELPgQsMLKTGDRQWFYFTPRSRKYPNAARSSRGTETGYWKATGKDRVIEY-NSRSVGL 117
69***************************.99**************96347888999************************************99.999**** PP
NAM 103 kktLvfykgrapkgektdWvmheyrl 128
kktLvfy+grap+ge+tdWvmhey++
336483 118 KKTLVFYRGRAPNGERTDWVMHEYTM 143
************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51005 | 52.959 | 16 | 166 | IPR003441 | NAC domain |
| SuperFamily | SSF101941 | 1.7E-55 | 17 | 166 | IPR003441 | NAC domain |
| Pfam | PF02365 | 3.1E-25 | 18 | 142 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0070301 | Biological Process | cellular response to hydrogen peroxide | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005789 | Cellular Component | endoplasmic reticulum membrane | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 548 aa Download sequence Send to blast |
MADSSPDSCF KGGKFSAPGF RFHPTDEELV MYYLKRKICR KRLRVNVIGV VDVYKMDPEE 60 LPGQSMLKTG DRQWFYFTPR SRKYPNAARS SRGTETGYWK ATGKDRVIEY NSRSVGLKKT 120 LVFYRGRAPN GERTDWVMHE YTMDEDELGR CKNPQDYYAL YKLFKKSGAG PKNGEQYGAP 180 FQEEEWVDDD NEDENAIAVP EQPVVRYEDT RRVDGRRLFN PVSLQLEDID ELLNGIPNAP 240 GVPPRCIPQV NSEEELQSTL MNNSAREFLP NDQPYNRPSS FDSLETAEVT SAPKTSGIAP 300 LVFEKEDYIE MDDLLIPELG ASSTEKAVQL SNHGGFGDFD EFDQLFHDVS MSLDMEPIDQ 360 GLLRICLIRQ FLYQQFQDQT PENQLDNIMD PSTTLDQFTN DIWFQDDQAI LFDQQQSFSG 420 AFASPSSGVI PDSTNPTMSV NAQGQEHQNG GGPTSQFSSA LWALMDSIPS TPASACEGPL 480 NRTFVRMSSF SRMRFNGKAN VTPVTTTIAK KGIRNRGFLL LSIVGALCAI FWVFIATVRV 540 SGRNVFS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 4e-44 | 18 | 143 | 17 | 140 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator activated by proteolytic cleavage through regulated intramembrane proteolysis (RIP). Transcriptional activator that acts as positive regulator of AOX1A during mitochondrial dysfunction. Binds directly to AOX1A promoter. Mediates mitochondrial retrograde signaling. {ECO:0000269|PubMed:24045017}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00179 | DAP | Transfer from AT1G34190 | Download |
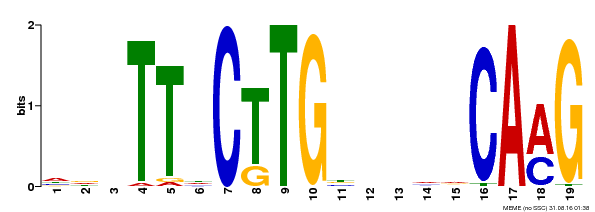
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 336483 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By cold and drought stresses. {ECO:0000269|PubMed:17158162}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF361813 | 0.0 | AF361813.1 Arabidopsis thaliana F23M19.13/F23M19.13 mRNA, complete cds. | |||
| GenBank | AY133552 | 0.0 | AY133552.1 Arabidopsis thaliana At1g34190/F23M19.13 mRNA, complete cds. | |||
| GenBank | BT000730 | 0.0 | BT000730.1 Arabidopsis thaliana clone RAFL09-58-M22 (R18947) putative NAM (no apical meristem) protein (At1g34190) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020868493.1 | 0.0 | LOW QUALITY PROTEIN: NAC domain-containing protein 17 | ||||
| Swissprot | Q9XIC5 | 0.0 | NAC17_ARATH; NAC domain-containing protein 17 | ||||
| TrEMBL | D7KJF5 | 0.0 | D7KJF5_ARALL; ANAC017 | ||||
| STRING | fgenesh1_pg.C_scaffold_1003048 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM2812 | 26 | 67 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G34190.1 | 0.0 | NAC domain containing protein 17 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 336483 |
| Entrez Gene | 9327144 |



