 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 327634 | ||||||||
| Common Name | ARALYDRAFT_327634 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; malvids; Brassicales; Brassicaceae; Camelineae; Arabidopsis
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 412aa MW: 47211.4 Da PI: 6.0303 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 136.7 | 1.5e-42 | 6 | 135 | 3 | 128 |
NAM 3 pGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp..kkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevlsk..kgelvg 101
+G+rF Pt eel+++yLk+k+ gk+ ++e+i+e++i++++P Lp +k+k+++ wyfF++++ ++a++k ++r+t+sgyWkatg d+++++k ++ ++g
327634 6 VGYRFYPTGEELINHYLKNKILGKTWLVDEAISEINICSYDPIYLPslSKIKSDDPVWYFFCPKEYTSAKKKVTKRTTSSGYWKATGVDRKIKDKrgNRGEIG 108
7************************999999***************5449999999**************************************9666667** PP
NAM 102 lkktLvfykgrapkgektdWvmheyrl 128
+kktLv+y+gr pkg+ t Wvmhey++
327634 109 IKKTLVYYEGRVPKGVWTPWVMHEYHI 135
*************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51005 | 45.577 | 4 | 153 | IPR003441 | NAC domain |
| SuperFamily | SSF101941 | 1.44E-46 | 5 | 153 | IPR003441 | NAC domain |
| Pfam | PF02365 | 6.1E-25 | 6 | 135 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005886 | Cellular Component | plasma membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 412 aa Download sequence Send to blast |
MVKDLVGYRF YPTGEELINH YLKNKILGKT WLVDEAISEI NICSYDPIYL PSLSKIKSDD 60 PVWYFFCPKE YTSAKKKVTK RTTSSGYWKA TGVDRKIKDK RGNRGEIGIK KTLVYYEGRV 120 PKGVWTPWVM HEYHITCLPQ DQRNYVICQV MYKGEDGDVP SGGNNSSEPS QSMVSDSNTV 180 RETITTAPEF EQPGQENFFG MSVDDLRTPM NEQKQEDFSL WDVLDPDMLF SDNNNSTVQP 240 QAPHLAPNDD EFLGGLRHVN REQVEYLFAN EDFISRPTLS VTENRNDHRP KKALSGIIVD 300 YSSDSNSDAE SISATSYQGT SSPGDDSVGS SNRHFLIHTD TSRLVLLWRR REAELVYYRR 360 SNGEKPQEST VYLSDEDDHR HHTLGGSHWQ HHIGFTNRQN LNPLMKFDRE R* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 7e-25 | 7 | 153 | 20 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 7e-25 | 7 | 153 | 20 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 7e-25 | 7 | 153 | 20 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 7e-25 | 7 | 153 | 20 | 165 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 7e-25 | 7 | 153 | 20 | 165 | NAC domain-containing protein 19 |
| 4dul_B | 7e-25 | 7 | 153 | 20 | 165 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator activated by proteolytic cleavage through regulated intramembrane proteolysis (RIP). Involved in salt stress response during seed germination and seedling growth. Binds the auxin-responsive IAA30 gene promoter and may serve as a molecular link that interconnects a developmental feedback loop of auxin signaling with a salt signal transduction pathway during seed germination (PubMed:21450938). {ECO:0000269|PubMed:21450938}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00426 | DAP | Transfer from AT4G01550 | Download |
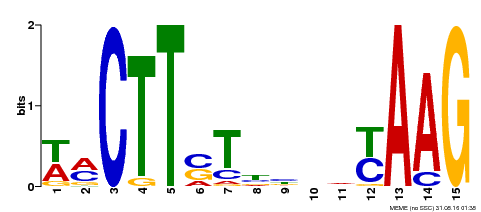
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 327634 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt stress and abscisic acid (ABA) in roots. {ECO:0000269|PubMed:21450938}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK229312 | 0.0 | AK229312.1 Arabidopsis thaliana mRNA for putative NAM-like protein, complete cds, clone: RAFL16-57-J19. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020876994.1 | 0.0 | LOW QUALITY PROTEIN: NAC domain-containing protein 69 | ||||
| Swissprot | Q9M126 | 0.0 | NAC69_ARATH; NAC domain-containing protein 69 | ||||
| TrEMBL | D7M4U1 | 0.0 | D7M4U1_ARALL; Uncharacterized protein | ||||
| STRING | fgenesh1_pm.C_scaffold_6002854 | 0.0 | (Arabidopsis lyrata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Malvids | OGEM1982 | 19 | 71 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G01550.1 | 0.0 | NAC domain containing protein 69 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 327634 |
| Entrez Gene | 9308955 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



