 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 30147.m013814 | ||||||||
| Common Name | RCOM_1507500 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Euphorbiaceae; Acalyphoideae; Acalypheae; Ricinus
|
||||||||
| Family | C2H2 | ||||||||
| Protein Properties | Length: 1736aa MW: 196560 Da PI: 8.126 | ||||||||
| Description | C2H2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | zf-C2H2 | 12.4 | 0.00048 | 1620 | 1645 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH.T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt.H 23
y+C C++sF +k +L +H r+ +
30147.m013814 1620 YQCDieGCTMSFGSKQELAVHKRNiC 1645
99********************9877 PP
| |||||||
| 2 | zf-C2H2 | 12.7 | 0.0004 | 1645 | 1667 | 3 | 23 |
ET..TTTEEESSHHHHHHHHHHT CS
zf-C2H2 3 Cp..dCgksFsrksnLkrHirtH 23
Cp Cgk+F ++ +L++H r+H
30147.m013814 1645 CPvkGCGKTFLSHKYLVQHRRVH 1667
9999*****************99 PP
| |||||||
| 3 | zf-C2H2 | 11.5 | 0.00091 | 1703 | 1729 | 1 | 23 |
EEET..TTTEEESSHHHHHHHHHH..T CS
zf-C2H2 1 ykCp..dCgksFsrksnLkrHirt..H 23
y+C Cg++F+ s++ rH r+ H
30147.m013814 1703 YVCAeeGCGQTFRFVSDFSRHKRKtgH 1729
89********************99666 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00545 | 4.7E-15 | 29 | 70 | IPR003349 | JmjN domain |
| PROSITE profile | PS51183 | 14.52 | 30 | 71 | IPR003349 | JmjN domain |
| Pfam | PF02375 | 1.9E-14 | 31 | 64 | IPR003349 | JmjN domain |
| PROSITE profile | PS51184 | 33.863 | 196 | 365 | IPR003347 | JmjC domain |
| SMART | SM00558 | 1.3E-49 | 196 | 365 | IPR003347 | JmjC domain |
| SuperFamily | SSF51197 | 4.12E-26 | 210 | 384 | No hit | No description |
| Pfam | PF02373 | 4.7E-37 | 229 | 348 | IPR003347 | JmjC domain |
| Gene3D | G3DSA:3.30.160.60 | 1.9E-4 | 1611 | 1642 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| SMART | SM00355 | 8.9 | 1620 | 1642 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE profile | PS50157 | 12.57 | 1643 | 1672 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 1.58E-6 | 1643 | 1679 | No hit | No description |
| SMART | SM00355 | 0.0033 | 1643 | 1667 | IPR015880 | Zinc finger, C2H2-like |
| Gene3D | G3DSA:3.30.160.60 | 1.4E-5 | 1644 | 1671 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE pattern | PS00028 | 0 | 1645 | 1667 | IPR007087 | Zinc finger, C2H2 |
| Gene3D | G3DSA:3.30.160.60 | 9.1E-8 | 1672 | 1697 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 10.533 | 1673 | 1702 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.0038 | 1673 | 1697 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1675 | 1697 | IPR007087 | Zinc finger, C2H2 |
| SuperFamily | SSF57667 | 3.35E-9 | 1683 | 1727 | No hit | No description |
| Gene3D | G3DSA:3.30.160.60 | 3.2E-9 | 1698 | 1726 | IPR013087 | Zinc finger C2H2-type/integrase DNA-binding domain |
| PROSITE profile | PS50157 | 11.032 | 1703 | 1734 | IPR007087 | Zinc finger, C2H2 |
| SMART | SM00355 | 0.85 | 1703 | 1729 | IPR015880 | Zinc finger, C2H2-like |
| PROSITE pattern | PS00028 | 0 | 1705 | 1729 | IPR007087 | Zinc finger, C2H2 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009826 | Biological Process | unidimensional cell growth | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0033169 | Biological Process | histone H3-K9 demethylation | ||||
| GO:0035067 | Biological Process | negative regulation of histone acetylation | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003676 | Molecular Function | nucleic acid binding | ||||
| GO:0046872 | Molecular Function | metal ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1736 aa Download sequence Send to blast |
MAAPSGLVTE PTASQQPQEV FQWLKNLPLA PEYHPTLAEF QDPIAYIFKI EKEASKYGIC 60 KIVPPVLAAP KKAAIANLNR SLAARSSSSK SAPTFTTRQQ QIGFCPRKPR PVQKPVWQSG 120 ENYTFQEFEA KAKSFEKSYF KKCPKKTAFS PLEVETLYWK ATVDKPFSVE YANDMPGSAF 180 SVKKMSGGKE IIEGVTVGET EWNMRGVSRA KGSLLRFMKE EIPGVTSPMV YVAMMFSWFA 240 WHVEDHDLHS LNYLHLGAGK TWYGVPKEAA VAFEEVVRDH GYGGEINPLV TFSVLGEKTT 300 VMSPEVFVTA GVPCCRLVQN AGEFVVTFPR AYHSGFSHGF NCGEAANIAT PEWLRVAKDA 360 AIRRASINYP PMVSHFQLLY DLALELCTRM PVSISAKPRS SRLKDKQKGE GETLVKEQFV 420 QNVIHNNELL HILGKGSSVV LLPRSSSDIS VCSDLQRNYG IDQSKGTISV KEKFASLCER 480 NRFSSLNGNE NKHTTNTRTE NKGTTHGDKL SDQRLFSCVT CGILSFDCIA VVQPTETAAR 540 YLMSADCSFF NDWIVGSGAT NNRLTTTNGD PNTCQLDQPT GWVENSVVDH LYDVPVQSVN 600 YQPQKIDKSK VNSNATMQGE SSALGLLALN YGNSSDSEED QDEPDVSDHA IDMPTCSSEN 660 KYKYQNCALP SFKQECHHDE TVSHTLSLVT LDCGDKVSLQ TDDCHKEHGD RAGNFKDGTP 720 DCFLDFGTDN MEPNGSECRF GDAVSISHIN SNCSPAVHDT EKMKFRRVVP RGNGDMPFAQ 780 RSDEDSSRMH VFCLEHAVEV EQQFRSIGGV HILLLCHPEY PRLEAEAKLV SEELGIDHLW 840 NDIAFRDATK NDEENIQSAL DSEEAIPGNG DWAVKLGINL FYSASLSHSS LYSKQMPYNS 900 VIYKAFGRVS PASSPTKLNV YGRRSGKQKK VVAGRWCGKV WMSNQVHNFL LKNASEDRDQ 960 EEEQDGSFHG WKMLDEKVER KLQNFYKTET ALAAAKSVRK RKLTTVTRPI KKVKSPETEA 1020 AASDDLEEDV SHKQHTKVYS RKQTKHIERE VSYDSLDNSH QQHGKTHRIK QAKSVERDDA 1080 VSDDSLGGNT HQQQHRRIPR SKQVKYIDSE NDVSDTLIGS NSQKQHSRIP KPRSKQVKYI 1140 RRKSEISDDS LEGDINEWHG RGPRSTQAGF YGREVAVSDD SLEESSHRPL GTVRRSKQAA 1200 YFESEDALSD DSVENSSLQQ NRRVSRGSLA KYFEREDEVS DDMLEENTFQ QCTGSHRSRK 1260 KKFVDREGEV SDNLLDESTQ LQHRRIFRSK QGRFVEREDE VSDDSLEDNS QQQRRIPTSK 1320 RAKFVESEDA VSDELEDDTH LKHWRIPRSK KAKIMEREAC SDDLQENDAQ WQPRKTPRGK 1380 PVKFIEREDV SDDLEEATHW QQRKTSRHKP VTSFEREDVS DDLQEENTRW QPRKTTRGKQ 1440 TKFFEKEDVS DDLQEDDTRW HPSKTPKSKQ AKYAELEDAV SDDLLEDNST KQHRRILRNK 1500 QKSGTLGKVN RESIQHVKQG TARAKKKENS KSIKKEKEVK QENPGFRNSK SGRLFESHVE 1560 EEVEGGPSTR LRKRPSKASK ESETKLKEKL QSNKKKVRGS ASAVKRASGQ KNNKDEDAEY 1620 QCDIEGCTMS FGSKQELAVH KRNICPVKGC GKTFLSHKYL VQHRRVHLDD RPLKCPWKGC 1680 KVTFKWAWAR TEHIRVHTGA RPYVCAEEGC GQTFRFVSDF SRHKRKTGHS VKKSRN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6ip0_A | 2e-78 | 21 | 513 | 6 | 457 | Transcription factor jumonji (Jmj) family protein |
| 6ip4_A | 2e-78 | 21 | 513 | 6 | 457 | Arabidopsis JMJ13 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 995 | 1001 | KSVRKRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Histone demethylase that demethylates 'Lys-27' (H3K27me) of histone H3. Demethylates both tri- (H3K27me3) and di-methylated (H3K27me2) H3K27me (PubMed:21642989, PubMed:27111035). Demethylates also H3K4me3/2 and H3K36me3/2 in an in vitro assay (PubMed:20711170). Involved in the transcriptional regulation of hundreds of genes regulating developmental patterning and responses to various stimuli (PubMed:18467490). Binds DNA via its four zinc fingers in a sequence-specific manner, 5'-CTCTGYTY-3', to promote the demethylation of H3K27me3 and the regulation of organ boundary formation (PubMed:27111034, PubMed:27111035). Involved in the regulation of flowering time by repressing FLOWERING LOCUS C (FLC) expression (PubMed:15377760). Interacts with the NF-Y complexe to regulate SOC1 (PubMed:25105952). Mediates the recruitment of BRM to its target loci (PubMed:27111034). {ECO:0000269|PubMed:15377760, ECO:0000269|PubMed:18467490, ECO:0000269|PubMed:20711170, ECO:0000269|PubMed:21642989, ECO:0000269|PubMed:25105952, ECO:0000269|PubMed:27111034, ECO:0000269|PubMed:27111035}. | |||||
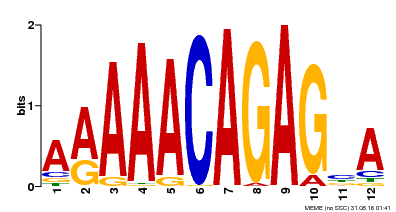
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00608 | ChIP-seq | Transfer from AT3G48430 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | 30147.m013814 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015580157.1 | 0.0 | lysine-specific demethylase REF6 | ||||
| Swissprot | Q9STM3 | 0.0 | REF6_ARATH; Lysine-specific demethylase REF6 | ||||
| TrEMBL | B9RAP0 | 0.0 | B9RAP0_RICCO; Nucleic acid binding protein, putative | ||||
| STRING | XP_002511265.1 | 0.0 | (Ricinus communis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF5648 | 32 | 50 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G48430.1 | 0.0 | relative of early flowering 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 30147.m013814 |



