 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | 26252 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Ostreococcus
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1094aa MW: 120634 Da PI: 6.6192 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 160.4 | 3.4e-50 | 48 | 165 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltl..elktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcywlLe 103
k+ k+rwl+n+e++++L n++ + ++ +++ rp g+l+L+nrk+vr+frkDG++w+kkkdgkt+rE+hekLKvg+ve+l+cyY+hsee+++fqrrcywlL+
26252 48 KTaKTRWLRNTEVCDVLLNYAAYGFEPsvDAPVRPLGGTLFLINRKVVRFFRKDGHNWQKKKDGKTIRETHEKLKVGTVELLNCYYTHSEEDAKFQRRCYWLLN 151
44499******************99987799************************************************************************* PP
CG-1 104 eelekivlvhylevk 118
++ e vlvhyl+vk
26252 152 MD-EGAVLVHYLTVK 165
**.********9885 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 70.169 | 43 | 170 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 1.1E-50 | 46 | 165 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 9.9E-46 | 50 | 163 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF01833 | 4.8E-5 | 475 | 549 | IPR002909 | IPT domain |
| SuperFamily | SSF81296 | 2.64E-14 | 475 | 560 | IPR014756 | Immunoglobulin E-set |
| Gene3D | G3DSA:2.60.40.10 | 5.5E-4 | 475 | 562 | IPR013783 | Immunoglobulin-like fold |
| Gene3D | G3DSA:1.25.40.20 | 1.2E-13 | 710 | 837 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.34E-13 | 735 | 834 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.37E-12 | 741 | 833 | No hit | No description |
| PROSITE profile | PS50297 | 14.186 | 744 | 825 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 9.404 | 777 | 809 | IPR002110 | Ankyrin repeat |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1094 aa Download sequence Send to blast |
MRRRVAGRAI ETIPTAPSGA TIPARALGEP TRAPTDAAAR ERANEVVKTA KTRWLRNTEV 60 CDVLLNYAAY GFEPSVDAPV RPLGGTLFLI NRKVVRFFRK DGHNWQKKKD GKTIRETHEK 120 LKVGTVELLN CYYTHSEEDA KFQRRCYWLL NMDEGAVLVH YLTVKKEPQR PSSGVATPGG 180 AARGALGSMG RDGKRAIGSK APLSSKIGNR ALSSKIGSVA SKIKSASTAG RRGAGRSRKD 240 VELESFLDGA ADDFSRQNVS DMLNRDDEDL ALIFQETGLD DAEQGASEWK YALDSEYISD 300 DDAPATSLSR LMREFNGLDD EYDDDCVDGR ARENSDQQTP TFSRSLVHSD SPRTGGEMSI 360 EEFERRIERI RSQWQNQVTG DGDSTSMERI ERQINTLEQD VERAMASSLQ RASDDPGGEP 420 STSAGDADAA DELSGHRANA TERGDAETSM HTRRTSGIVR HVPATPSGVH VLWSIVDFTP 480 SWDDVSGGAK VIITGNPLVE LEPGIGMCCV FGTIAVPVEQ LAPNVLKCYA PAHAPGVVSM 540 FLVMESGNGH PVSEISSFEF MESLDPSRGV DVDRRDMIDQ SANMSDRDFQ MRLVQLLTTL 600 GSDSSNSVGN DSGEKSGDRT THSSALVNDS VMHMNALSAL RAANRLELDP YNLDGVKNEE 660 LVVLLSGMLQ ARLKSVIVHE NRRMKARRAL PSSAVAMQEV EEVAKTGVIS DKIVETAVEK 720 TQQTHKALLK VAFTPSAYKR KDQTGLTLFH CCAALGIEWA VRAMCVTGVD LNHTDAYNRS 780 ALHWAVARGH EMVVATLLNY GAKSRSMCQW EGESFTPAEL AVRCGYEGIS AYISEANLAS 840 ALENINLRNS GIPRQRCRTS RGGGVFQAHR FRRTILSDTE GSDSEEHSVR RSTRVNSEKK 900 RAYHRRKTSR LTASTTADGD DSETEAAFVV KRAENARRSL LDTLYNLNVK PEFIDADIHA 960 RLGKRRGRRV RQVDVQSLMS ELLTPVGDNP THSEMMSPSR RPVLRARRAP KPVVGVSITN 1020 DVKGREDSED SESLDDDEDE EAIQKNFSRI QTSLQSAHSR TQYLRLRRLT NQLRVELKKL 1080 KDDGQFSDDD DDR* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 680 | 687 | NRRMKARR |
| 2 | 960 | 969 | RLGKRRGRRV |
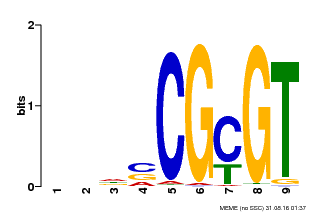
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CP000590 | 0.0 | CP000590.1 Ostreococcus lucimarinus CCE9901 chromosome 10, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_001420194.1 | 0.0 | predicted protein | ||||
| TrEMBL | A4S420 | 0.0 | A4S420_OSTLU; Uncharacterized protein | ||||
| STRING | ABO98487 | 0.0 | (Ostreococcus 'lucimarinus') | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP5132 | 8 | 9 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G09410.2 | 1e-41 | ethylene induced calmodulin binding protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | 26252 |



