 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc07122.1.g00010.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 247aa MW: 26299.7 Da PI: 5.9184 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 56.9 | 5e-18 | 2 | 50 | 3 | 55 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+ GVr+++ +gr++AeIrdp++ + r++lg+f+ta+eAa a+++a+++++g
Zpz_sc07122.1.g00010.1.am.mk 2 FLGVRRRP-WGRYAAEIRDPTT---KERHWLGTFDTAQEAALAYDRAALSMKG 50
56******.**********965...5************************998 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 5.29E-26 | 1 | 57 | No hit | No description |
| PROSITE profile | PS51032 | 20.547 | 1 | 58 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 7.5E-30 | 1 | 64 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.75E-20 | 2 | 59 | IPR016177 | DNA-binding domain |
| Gene3D | G3DSA:3.30.730.10 | 3.1E-28 | 2 | 58 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.4E-8 | 2 | 13 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 1.8E-11 | 4 | 50 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 7.4E-8 | 24 | 40 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010102 | Biological Process | lateral root morphogenesis | ||||
| GO:0010432 | Biological Process | bract development | ||||
| GO:0010451 | Biological Process | floral meristem growth | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 247 aa Download sequence Send to blast |
RFLGVRRRPW GRYAAEIRDP TTKERHWLGT FDTAQEAALA YDRAALSMKG AQARTNFVYS 60 HTAYNYPPFL APFHAQPSYG YAQSSMHQYG GQHGMGASSH IGSYHPHYQA PAGSGASSAS 120 GECSMPVAVD RADGSLLDGG HNFLFSSADD NSGYLSSVVP ESCLRPRSSA AAAEDMRRYS 180 DADAYGMMGL REDVDDLAQM VAGFWGDAGD AEQLGACAFP AGGGHGGDIV SSSQGSDAYS 240 PFSFLSH |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 2e-17 | 2 | 57 | 6 | 62 | ATERF1 |
| 3gcc_A | 2e-17 | 2 | 57 | 6 | 62 | ATERF1 |
| 5wx9_A | 2e-17 | 1 | 62 | 14 | 76 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Required to prevent the formation of axillary meristems within the spikelet meristem and permit the subsequent establishment of floral meristem identity (PubMed:12835399, PubMed:14503923). Mediates the transition from spikelet to floret meristem (PubMed:14503923). Determines the transition from panicle branching to spikelet formation. May specify floral organ identity by regulating the class B genes (Agamous-like genes) MADS6 and MADS17, as well as class E genes MADS1, MADS7 and MADS8 in floral meristem (PubMed:26744119). Possesses transactivation activity (PubMed:12835399). {ECO:0000269|PubMed:12835399, ECO:0000269|PubMed:14503923, ECO:0000269|PubMed:26744119}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00520 | DAP | Transfer from AT5G18560 | Download |
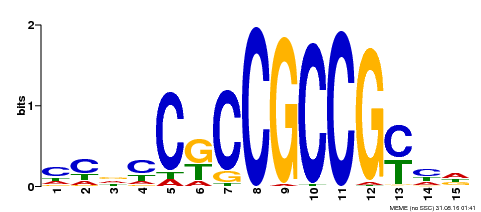
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc07122.1.g00010.1.am.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT036365 | 1e-101 | BT036365.1 Zea mays full-length cDNA clone ZM_BFb0108M21 mRNA, complete cds. | |||
| GenBank | KJ727547 | 1e-101 | KJ727547.1 Zea mays clone pUT5407 AP2-EREBP transcription factor (EREB151) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025804267.1 | 1e-120 | ethylene-responsive transcription factor FZP | ||||
| Swissprot | Q8H3Q1 | 1e-112 | FZP_ORYSJ; Ethylene-responsive transcription factor FZP | ||||
| TrEMBL | A0A1E5UMV0 | 1e-123 | A0A1E5UMV0_9POAL; Ethylene-responsive transcription factor ERF086 | ||||
| STRING | Pavir.Ba00192.1.p | 1e-117 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP10525 | 28 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G18560.1 | 3e-41 | ERF family protein | ||||



