 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc03663.1.g00040.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 697aa MW: 77307.3 Da PI: 9.4452 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 25.3 | 2.7e-08 | 482 | 520 | 17 | 55 |
HHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 17 elFeknrypsaeereeLAkklgLterqVkvWFqNrRake 55
e+ +k +yps++++ LA+++gL+ +q+ +WF N+R ++
Zpz_sc03663.1.g00040.1.am.mk 482 EMHYKWPYPSETQKGALAESTGLDLKQINNWFINQRKRH 520
4555789*****************************885 PP
| |||||||
| 2 | ELK | 36 | 1.5e-12 | 440 | 461 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELKh+Ll+KYsgyL+sLkqE+s
Zpz_sc03663.1.g00040.1.am.mk 440 ELKHHLLKKYSGYLSSLKQELS 461
9*******************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 8.4E-18 | 274 | 321 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 2.8E-20 | 275 | 319 | IPR005540 | KNOX1 |
| SMART | SM01256 | 2.7E-24 | 328 | 388 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 2.5E-16 | 334 | 386 | IPR005541 | KNOX2 |
| Pfam | PF03789 | 1.8E-9 | 440 | 461 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 11.331 | 440 | 460 | IPR005539 | ELK domain |
| SMART | SM01188 | 4.9E-7 | 440 | 461 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.584 | 460 | 523 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 6.84E-19 | 461 | 536 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 2.2E-12 | 462 | 527 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.8E-27 | 465 | 525 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 3.51E-11 | 472 | 524 | No hit | No description |
| Pfam | PF05920 | 4.9E-16 | 480 | 519 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 498 | 521 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001708 | Biological Process | cell fate specification | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 697 aa Download sequence Send to blast |
MARRAWGMID LRLFNSGSLP FPRRDTNQQE RVRGRSQSLS EPRAPHIPRR PARSPPHARA 60 FPLPAHVLSS QGYCQRHYHR NNSLGRLELR GYYHHRQGYI LSRSLHCAPR PAPGFRREKA 120 KLQAPPPPSP LLSPKPFSSF SPPENFTSTP NHPLFSAGSR SSDNGRRARS LHQYIPGDRP 180 TPMEEITHHF GVGGSGQGQQ QHHHSWVSAL GAVLAPPPSA GLPLTLNTVA AGNSGAHSGS 240 SNPVLQLANG GSLLDACVKA KQPSSSSPYA GDVEAIKAKI ISHPHYYSLL AAYLECQKAS 300 PVGAPPEVSA RLTAIAQEVE ARQRTSLVGL GSATEPELDQ FMVSHFYRLI NEAYHEMLVK 360 FREELTRPLQ EAMEFMRRVE SQLNSLSISG RSLRRILSSD ALVHYAYPAQ RGTGSSEEDQ 420 EGSGGETELP EVDAHGVDQE LKHHLLKKYS GYLSSLKQEL SKKKKKGKLP KEARQQLLNW 480 WEMHYKWPYP SETQKGALAE STGLDLKQIN NWFINQRKRH WKPSEEMHHL MMDGYHTPNA 540 FYMDGHFLNE PGLYRFDRTC DAIDAPDAPR CGEVRIARSM HGLPLWVGIL EEAAVAKLVD 600 WTGRVPNAAL CCDLRSCGLG SLAPGRAMEL PAAVSRSPSR LQARVVQDSC PESRHDAEAD 660 AGPPPRRSPA FYSSVFAQIE EIGWERLVSS KGNDGIS |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to RNA (PubMed:17965274). Possible transcription factor that regulates genes involved in development. Mutations in KN-1 alter leaf development. Foci of cells along the lateral vein do not differentiate properly but continue to divide, forming knots. May participate in the switch from indeterminate to determinate cell fates. Probably binds to the DNA sequence 5'-TGAC-3'. {ECO:0000269|PubMed:17965274}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | Transfer from GRMZM2G017087 | Download |
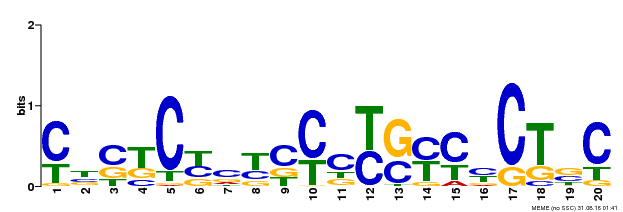
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc03663.1.g00040.1.am.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | DQ317421 | 1e-139 | DQ317421.1 Chasmanthium latifolium KNOTTED1 homeodomain protein (KN1) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004981913.1 | 0.0 | homeotic protein knotted-1 | ||||
| Swissprot | P24345 | 0.0 | KN1_MAIZE; Homeotic protein knotted-1 | ||||
| TrEMBL | A0A0A9CTZ3 | 0.0 | A0A0A9CTZ3_ARUDO; Kn1 | ||||
| STRING | GRMZM2G017087_P02 | 0.0 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9886 | 31 | 45 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 3e-78 | KNOTTED-like from Arabidopsis thaliana | ||||



