 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc03228.1.g00010.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 608aa MW: 64657 Da PI: 10.0287 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 59.9 | 5.9e-19 | 432 | 482 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
++++GVr ++ +gr++AeIrdp k++r++lg+f++ae+Aa+a++aa++ l+g
Zpz_sc03228.1.g00010.1.am.mk 432 TRFRGVRKRP-WGRYAAEIRDPA---KKARVWLGTFDSAEDAARAYDAAARTLRG 482
589*******.**********83...36*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50929 | 10.674 | 1 | 44 | IPR011527 | ABC transporter type 1, transmembrane domain |
| Pfam | PF00664 | 4.2E-7 | 1 | 44 | IPR011527 | ABC transporter type 1, transmembrane domain |
| SuperFamily | SSF90123 | 8.24E-5 | 2 | 49 | IPR011527 | ABC transporter type 1, transmembrane domain |
| Gene3D | G3DSA:1.20.1560.10 | 2.3E-4 | 2 | 46 | No hit | No description |
| CDD | cd00018 | 7.60E-17 | 432 | 490 | No hit | No description |
| Pfam | PF00847 | 2.4E-12 | 432 | 482 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.31E-21 | 433 | 491 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 1.3E-36 | 433 | 496 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 23.472 | 433 | 490 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 7.9E-31 | 433 | 490 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.9E-9 | 434 | 445 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.9E-9 | 456 | 472 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0055085 | Biological Process | transmembrane transport | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005524 | Molecular Function | ATP binding | ||||
| GO:0042626 | Molecular Function | ATPase activity, coupled to transmembrane movement of substances | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 608 aa Download sequence Send to blast |
MGEHLTKRIR EQMLAKILTF EIGWFDRDEN SSGVICSQLA KDANICTELR RYLIIPLPAR 60 DDPTPRRETG RVVLCGGDGT AALTRAREVG TYRELFGCRL VLHCTAASVR SSRRGRRGTP 120 RTPVACGEKP ANLGDAFEGI SNTLSGARYL GTVGPAWTTQ AKQGIICAGR RLIIPFHLAY 180 GRKELGGATA PPPARNVCPC VRSIGTALPR AHEASSSSPT HHARHHPSTT GGDPAGRSSL 240 LSTPCQDACP GILPAGVTGK THQPHFLNSV RESRETRRSV SGPGFLLCCT HGAQRRQRAA 300 APMRCDACMI AAHEANGGGG AGAHGAPDQS PSPHKPERAG TPGSPHRRQK KKRREPRQGG 360 WWAGGGGSVA GGWECAVRGG SANKQTFQLS PPQHPRQFPD SPAPRTAGFI GGEGMRRGGP 420 VAAAGPEGDL VTRFRGVRKR PWGRYAAEIR DPAKKARVWL GTFDSAEDAA RAYDAAARTL 480 RGPKARTNFP LPVAAHHHHQ RHAAYTAPYT AVAARPAISS MSSTVESFGG ARSRPALPPR 540 PPPPPIPDGD CRSDCGSSAS VVDDDCADAA TSPSCRLPLA LDLNLPPGDC GFVCYGDEDE 600 LRLTELRL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 2e-20 | 434 | 493 | 6 | 66 | ATERF1 |
| 3gcc_A | 2e-20 | 434 | 493 | 6 | 66 | ATERF1 |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 346 | 354 | RRQKKKRRE |
| 2 | 349 | 353 | KKKRR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00194 | DAP | Transfer from AT1G50640 | Download |
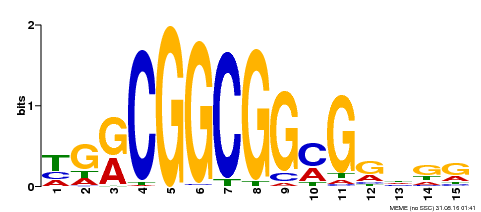
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc03228.1.g00010.1.am.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK359709 | 1e-65 | AK359709.1 Hordeum vulgare subsp. vulgare mRNA for predicted protein, complete cds, clone: NIASHv1101F03. | |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5123 | 27 | 58 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G50640.1 | 9e-21 | ethylene responsive element binding factor 3 | ||||



