 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc01720.1.g00010.1.sm.mkhc | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 890aa MW: 97091.6 Da PI: 5.8708 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130.7 | 5.1e-41 | 153 | 230 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+CqvegC adl+ ak+yhrrhkvCe+h+ka++++v ++ qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+q+
Zpz_sc01720.1.g00010.1.sm.mkhc 153 ACQVEGCVADLTAAKDYHRRHKVCEMHAKASTAVVGKTVQRFCQQCSRFHLLEEFDEGKRSCRRRLAGHNRRRRKTQP 230
5**************************************************************************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.1E-33 | 148 | 215 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.223 | 151 | 228 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 5.76E-38 | 152 | 232 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 2.6E-29 | 154 | 227 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 3.11E-8 | 668 | 775 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 9.649 | 670 | 737 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 5.3E-8 | 671 | 773 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 4.49E-9 | 675 | 775 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 890 aa Download sequence Send to blast |
MEAARLGAQS SQLYGGGLGE LDLNMREKRV FSWDLNDWSW DSEHFVATPV PASVTNGLEL 60 NSSPSSSEKA EAEVARNGAM RGDSDKRKRV VVIDECEAED QDLIENSGGG GALSLKIGSD 120 AVSPRTLESS GVNEEVRNGK KIKAQGGISS GPACQVEGCV ADLTAAKDYH RRHKVCEMHA 180 KASTAVVGKT VQRFCQQCSR FHLLEEFDEG KRSCRRRLAG HNRRRRKTQP DISFGGAASI 240 EDKELLSNLL KNLGSVAKSL ELKELSKLLE ACQSLQNGSN AGTSGTANAL ANNSAAEAAG 300 PSNSNLLFMN GSQCGQASSS AIPVQQKATM VANPEPRACK IKDFDLNDTC DYIEGFQNGQ 360 EGSPTPAFKA ADSPNCPSWM HQNSTQIPPQ TSGNSDSTST QSLSSSNGDA QLSDNLSSHL 420 EKLLNSSSDN FWASGLVFVM VRHQLAFMYN GQIMLDKRLA PSSFHYCKIL SVRPVAVPYS 480 AAVNFRIEGF NLVSASSRLI CSFEGCCIFQ EDTAAITDDA ELDDRETECL SFCCPAPGSR 540 GRGFIEVEDS GFSNSFFPFI IAEQDVCSEL SELESIFESF SHQPADVDDV ARNQALEFLN 600 ELGWLLHRAN IISKQEKMEP PLATFNLWRF RNLGIFAMER EWCAVIKMLL DFLFTGLVDV 660 GSQSPEEVVL SENLLHTAVQ RKSVQMVRFL LRYKANKSSK GTTQKNLFRP DDIGPSAITP 720 LHIAASTSDA EDVLDALTDD PGLVGISAWK NARDNTGFTP EDYARQRCND AYLNLVQKKI 780 DKHISEGHVV VGVPSSMCPV ITDFVKPGKV SLEIGRSVSM AAPVAKCHLC RNQALKYQSA 840 AVRTFLYRPA MLTMMGVAVV CVCVGILLHT MPRVYAAPTF RWELLERGAM |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 5e-30 | 147 | 227 | 4 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3'. {ECO:0000250}. | |||||
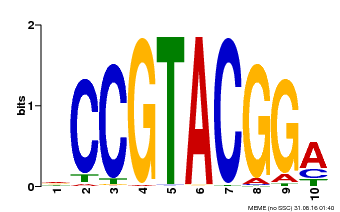
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00603 | PBM | Transfer from AT2G47070 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc01720.1.g00010.1.sm.mkhc |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004981153.1 | 0.0 | squamosa promoter-binding-like protein 6 | ||||
| Swissprot | Q75LH6 | 0.0 | SPL6_ORYSJ; Squamosa promoter-binding-like protein 6 | ||||
| TrEMBL | A0A0A9REY0 | 0.0 | A0A0A9REY0_ARUDO; Uncharacterized protein | ||||
| STRING | Si034095m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1998 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 1e-105 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



