 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc01397.1.g00150.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 229aa MW: 23602.4 Da PI: 10.5338 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 128.6 | 6.4e-40 | 27 | 124 | 3 | 98 |
TCP 3 gkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgtssssasec..eaesssss 81
++k +++++h+kv+gR+RR+R+++ caar+F+L++eLG+++d++tieWLl+qa+p+i+++tgt++++as++ +++s+sss
Zpz_sc01397.1.g00150.1.am.mk 27 VRKAPSKDRHSKVDGRGRRIRMPIICAARVFQLTRELGHKSDGQTIEWLLRQAEPSIIAATGTGTTPASFSttSPSSRSSS 107
689************************************************************999998888888888888 PP
TCP 82 asnsssg..kaaksaaksk 98
+++++++ +aa++++
Zpz_sc01397.1.g00150.1.am.mk 108 QQAPQIQlpSAAPAHH--V 124
8888888553333333..2 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51369 | 28.209 | 32 | 86 | IPR017887 | Transcription factor TCP subgroup |
| Pfam | PF03634 | 8.8E-34 | 32 | 111 | IPR005333 | Transcription factor, TCP |
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 229 aa Download sequence Send to blast |
MTTRPADGGG GGGDKQLVPA SNNAHGVRKA PSKDRHSKVD GRGRRIRMPI ICAARVFQLT 60 RELGHKSDGQ TIEWLLRQAE PSIIAATGTG TTPASFSTTS PSSRSSSQQA PQIQLPSAAP 120 AHHVLPHAAA AAPFILGKRV RGANDDTEVT AVGPAAGFWA LPARADFGQL WSFAAAPDMM 180 VAAVPEEASA ARYLPVAQAQ GNLNLLASFS GGPAGNGATA ARAEEESAR |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00524 | DAP | Transfer from AT5G23280 | Download |
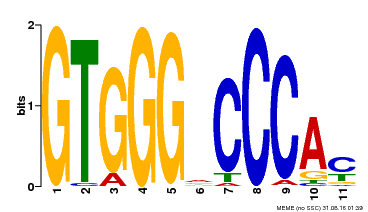
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc01397.1.g00150.1.am.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK240798 | 1e-103 | AK240798.1 Oryza sativa Japonica Group cDNA, clone: J065002G04, full insert sequence. | |||
| GenBank | AP004776 | 1e-103 | AP004776.3 Oryza sativa Japonica Group genomic DNA, chromosome 2, PAC clone:P0452F04. | |||
| GenBank | AP014958 | 1e-103 | AP014958.1 Oryza sativa Japonica Group DNA, chromosome 2, cultivar: Nipponbare, complete sequence. | |||
| GenBank | CP012610 | 1e-103 | CP012610.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 2 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025814152.1 | 4e-85 | transcription factor TCP7-like | ||||
| Swissprot | Q9FMX2 | 2e-66 | TCP7_ARATH; Transcription factor TCP7 | ||||
| TrEMBL | A0A0A9JXS1 | 8e-86 | A0A0A9JXS1_ARUDO; Uncharacterized protein | ||||
| STRING | OMERI02G35320.1 | 4e-80 | (Oryza meridionalis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1941 | 33 | 100 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G23280.1 | 1e-44 | TCP family protein | ||||



