 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc01350.1.g00230.1.sm.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 379aa MW: 41075.5 Da PI: 10.1512 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 39.6 | 1.3e-12 | 61 | 109 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ r +r+++g+f ++eAa+a++ a+++++g
Zpz_sc01350.1.g00230.1.sm.mk 61 SKYKGVVPQP-NGRWGAQIYE------RhQRVWIGTFTGEDEAARAYDVAAQRFRG 109
89****9888.8*********......44**********99*************98 PP
| |||||||
| 2 | B3 | 92.5 | 3.2e-29 | 193 | 298 | 1 | 90 |
EEEE-..-HHHHTT-EE--HHH.HTT..................---..--SEEEEEETTS-EEEEEE..EEETTEEEE-T CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh..................ggkkeesktltledesgrsWevkliyrkksgryvltk 63
f+k+ tpsdv+k++rlv+pk++ae+h +g + +++ l +ed sg++W+++++y+++s++yvlt+
Zpz_sc01350.1.g00230.1.sm.mk 193 FDKTVTPSDVGKLNRLVIPKQHAEKHfplhlpapaaaavdddvvSG-ECKGVLLNFEDVSGKVWRFRYSYWNSSQSYVLTR 272
89***************************99988888888777522.34899***************************** PP
THHHHHHHHT--TT-EEEEEE-SSSEE CS
B3 64 GWkeFvkangLkegDfvvFkldgrsef 90
GW++Fvk++gL +gD v F++ ++
Zpz_sc01350.1.g00230.1.sm.mk 273 GWSRFVKEKGLHAGDAVGFYRRA-AGK 298
********************533.444 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 2.22E-16 | 61 | 117 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 2.65E-23 | 61 | 117 | No hit | No description |
| Pfam | PF00847 | 1.8E-8 | 61 | 109 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 4.8E-19 | 62 | 117 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 9.2E-25 | 62 | 123 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.888 | 62 | 117 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 2.7E-38 | 188 | 313 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 8.86E-28 | 191 | 302 | No hit | No description |
| SuperFamily | SSF101936 | 5.76E-29 | 191 | 298 | IPR015300 | DNA-binding pseudobarrel domain |
| PROSITE profile | PS50863 | 12.337 | 193 | 308 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 4.3E-26 | 193 | 300 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 2.4E-20 | 193 | 308 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 379 aa Download sequence Send to blast |
MDSTSCLLDS ASSGASTGKK APAELIRTLQ RVGSGASAPG AEADSGGARR AAAAGGKLPS 60 SKYKGVVPQP NGRWGAQIYE RHQRVWIGTF TGEDEAARAY DVAAQRFRGR DAVTNFRPLA 120 DSDPDDAAEL RFLAARTKVE VVDMLRKHTY LDELAQNRRA FAAVAPPPSP STNGPADSSP 180 HPQPPSAARE HLFDKTVTPS DVGKLNRLVI PKQHAEKHFP LHLPAPAAAA VDDDVVSGEC 240 KGVLLNFEDV SGKVWRFRYS YWNSSQSYVL TRGWSRFVKE KGLHAGDAVG FYRRAAGKQL 300 YIDCKVRSRK TSPPGPVKAV RLFGVDLLTS TPSPASAAAT LQEEMTTMAM KNKRPRDFIV 360 TSSTPHMFFK KQCVDLALT |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 4e-47 | 190 | 308 | 11 | 118 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00025 | PBM | Transfer from AT1G68840 | Download |
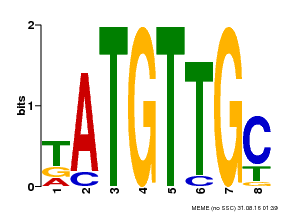
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc01350.1.g00230.1.sm.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT041962 | 0.0 | BT041962.2 Zea mays full-length cDNA clone ZM_BFb0089O18 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025807648.1 | 0.0 | AP2/ERF and B3 domain-containing protein Os05g0549800 | ||||
| Swissprot | Q6L4H4 | 1e-166 | Y5498_ORYSJ; AP2/ERF and B3 domain-containing protein Os05g0549800 | ||||
| TrEMBL | A0A3L6SXE9 | 0.0 | A0A3L6SXE9_PANMI; AP2/ERF and B3 domain-containing protein | ||||
| STRING | Si024576m | 0.0 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP456 | 38 | 203 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G68840.2 | 1e-106 | related to ABI3/VP1 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



