 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc00237.1.g00090.1.sm.mkhc | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 328aa MW: 35725.6 Da PI: 5.1638 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 58.6 | 1.5e-18 | 109 | 158 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkr.krfslgkfgtaeeAakaaiaarkkleg 55
+++G+r+++ +g+W+AeIrdp + +r++lg+f tae+Aa+a++ +++l+g
Zpz_sc00237.1.g00090.1.sm.mkhc 109 HFRGIRQRP-WGKWAAEIRDPH----KgTRVWLGTFNTAEDAARAYDVEARRLRG 158
69*******.**********72....35*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.730.10 | 7.5E-32 | 108 | 166 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 3.2E-11 | 109 | 158 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 23.42 | 109 | 166 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 8.37E-29 | 109 | 168 | No hit | No description |
| SMART | SM00380 | 1.7E-35 | 109 | 172 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.11E-21 | 109 | 167 | IPR016177 | DNA-binding domain |
| PRINTS | PR00367 | 2.8E-11 | 110 | 121 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 2.8E-11 | 132 | 148 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0034059 | Biological Process | response to anoxia | ||||
| GO:0071456 | Biological Process | cellular response to hypoxia | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 328 aa Download sequence Send to blast |
MCGGAILAEL IPPTRRAASR PVTAGGNLWP ASSENGGRGR DNKKHVAEID DFEAAFDEFD 60 DDFDEVEEDG DYGYDGSRAR PFVFASKPAF SSAHDGRAAR QRKRGSRRHF RGIRQRPWGK 120 WAAEIRDPHK GTRVWLGTFN TAEDAARAYD VEARRLRGSK AKVNFPAAGA MPRRAKARPA 180 PKAQPPPAAA QLASLGGQKK KQEELVAEPE TMMAAFNMDN LLDLTFPAAP PVTASSFTDS 240 ARSESGAPVK KQRTDHSSDG SATLELTDEL EFDPFMFFQL PYMDGGYESI DSLFAGGEAG 300 QDVNTVNSDM NGAGLWSFDE FPIDGAIF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 4e-22 | 108 | 166 | 4 | 63 | ATERF1 |
| 3gcc_A | 4e-22 | 108 | 166 | 4 | 63 | ATERF1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcriptional activator that may be involved in defense signaling pathway. Binds in vitro to the DNA sequence 5'-AGCCGCC-3' of the GCC-box element found in pathogenesis-related (PR) gene promoters. The transcriptional activation is enhanced in vitro by the presence of MPK12/BWMK1. {ECO:0000269|PubMed:12913152}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00230 | DAP | Transfer from AT1G72360 | Download |
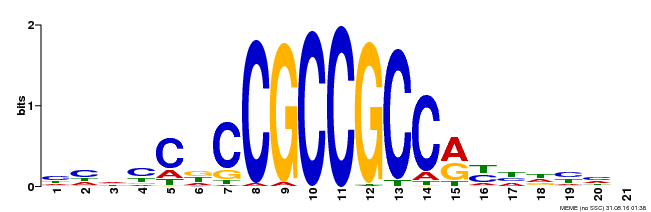
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc00237.1.g00090.1.sm.mkhc |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT034434 | 3e-81 | BT034434.1 Zea mays full-length cDNA clone ZM_BFc0170J03 mRNA, complete cds. | |||
| GenBank | BT039300 | 3e-81 | BT039300.1 Zea mays full-length cDNA clone ZM_BFc0004M21 mRNA, complete cds. | |||
| GenBank | BT063174 | 3e-81 | BT063174.1 Zea mays full-length cDNA clone ZM_BFc0035E09 mRNA, complete cds. | |||
| GenBank | EU941438 | 3e-81 | EU941438.1 Zea mays clone 1452173 mRNA sequence. | |||
| GenBank | EU970244 | 3e-81 | EU970244.1 Zea mays clone 342610 root abundant factor mRNA, complete cds. | |||
| GenBank | KJ726938 | 3e-81 | KJ726938.1 Zea mays clone pUT3484 AP2-EREBP transcription factor (EREB180) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025791485.1 | 1e-118 | ethylene-responsive transcription factor 1-like | ||||
| Refseq | XP_025791486.1 | 1e-118 | ethylene-responsive transcription factor 1-like | ||||
| Swissprot | Q6K7E6 | 5e-32 | ERF1_ORYSJ; Ethylene-responsive transcription factor 1 | ||||
| TrEMBL | A0A0A9VK36 | 1e-130 | A0A0A9VK36_ARUDO; Uncharacterized protein | ||||
| STRING | Pavir.Ib00526.1.p | 1e-125 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7987 | 30 | 40 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G72360.2 | 1e-28 | ERF family protein | ||||



