 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc00113.1.g00680.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 311aa MW: 33447.3 Da PI: 5.5391 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 65.3 | 1.3e-20 | 135 | 186 | 2 | 56 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkklege 56
+y+GVr+++ +g+++AeIrdp+++g r++lg+++t+ eAa+a+++a+ +++g+
Zpz_sc00113.1.g00680.1.am.mk 135 KYRGVRQRP-WGKYAAEIRDPKKRG--SRVWLGTYDTPVEAARAYDRAAFRMRGA 186
8********.***********8876..**************************95 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.730.10 | 1.8E-32 | 134 | 194 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 24.078 | 135 | 193 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 3.89E-32 | 135 | 194 | No hit | No description |
| SMART | SM00380 | 3.6E-39 | 135 | 199 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.05E-22 | 135 | 194 | IPR016177 | DNA-binding domain |
| Pfam | PF00847 | 1.9E-14 | 135 | 185 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.5E-13 | 136 | 147 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.5E-13 | 175 | 195 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 311 aa Download sequence Send to blast |
MDFSGEIDDF WLQNICEQFH DGEACLPVPE VYSASSTVAH PIPAPQHDFQ YAVFMPQQQV 60 ALVDLTQGYG DATAAAALSA KPAAPLIIQF GGEPSPRPSL TISLPPTTHV WGADVAPLPV 120 HSAAAASVDV EDLRKYRGVR QRPWGKYAAE IRDPKKRGSR VWLGTYDTPV EAARAYDRAA 180 FRMRGAKAIL NFPNEVGSGG ADFLTPQQPA SRSHSSKRKL HDAHQPDDVE AETAAKSIKT 240 EAFTSPASSQ TSLSPATTMA STVTSSSSSE AEMFPISPSG WSWEQFESVF GSLSPLSPHP 300 GHMGFPEVTV N |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 9e-28 | 134 | 199 | 4 | 69 | ATERF1 |
| 3gcc_A | 9e-28 | 134 | 199 | 4 | 69 | ATERF1 |
| Search in ModeBase | ||||||
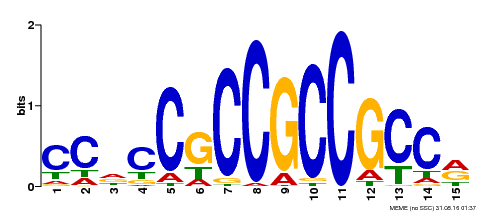
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00549 | DAP | Transfer from AT5G47230 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc00113.1.g00680.1.am.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AJ307662 | 4e-64 | AJ307662.1 Oryza sativa genomic DNA fragment, chromosome 2. | |||
| GenBank | AP004676 | 4e-64 | AP004676.3 Oryza sativa Japonica Group genomic DNA, chromosome 2, BAC clone:OJ1003_B06. | |||
| GenBank | AP005006 | 4e-64 | AP005006.3 Oryza sativa Japonica Group genomic DNA, chromosome 2, PAC clone:P0519E06. | |||
| GenBank | AP014958 | 4e-64 | AP014958.1 Oryza sativa Japonica Group DNA, chromosome 2, cultivar: Nipponbare, complete sequence. | |||
| GenBank | AY339378 | 4e-64 | AY339378.1 Oryza sativa (japonica cultivar-group) Ap25 mRNA, complete cds. | |||
| GenBank | CP012610 | 4e-64 | CP012610.1 Oryza sativa Indica Group cultivar RP Bio-226 chromosome 2 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025828794.1 | 1e-109 | ethylene-responsive transcription factor ERF105-like | ||||
| TrEMBL | A0A3L6PMC1 | 1e-108 | A0A3L6PMC1_PANMI; Putative AP2/EREBP transcription factor superfamily protein | ||||
| STRING | GRMZM2G057386_P01 | 1e-104 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP103 | 37 | 465 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G47230.1 | 2e-37 | ethylene responsive element binding factor 5 | ||||



