 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc00017.1.g00700.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 974aa MW: 104511 Da PI: 9.7582 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.1 | 1.7e-40 | 182 | 258 | 2 | 78 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
C v+gC+adls+++eyhrrhkvCe+hsk+pvv+v+g+e rfCqqCsrfh+l efDe+krsCr+rL++hn+rrrk+q+
Zpz_sc00017.1.g00700.1.am.mk 182 CAVDGCDADLSKCREYHRRHKVCEAHSKTPVVVVAGREMRFCQQCSRFHSLVEFDEAKRSCRKRLDGHNRRRRKPQP 258
**************************************************************************987 PP
| |||||||
| 2 | SRF-TF | 93.7 | 8.4e-30 | 897 | 947 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
k+ienk+nrqvtf+kRrng+lKKA+ELSvLCdaeva+iifs++gkl+e++s
Zpz_sc00017.1.g00700.1.am.mk 897 KQIENKINRQVTFAKRRNGLLKKAYELSVLCDAEVALIIFSNRGKLFEFCS 947
68***********************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.4E-34 | 174 | 243 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.129 | 179 | 256 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 2.22E-37 | 180 | 261 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.3E-30 | 182 | 255 | IPR004333 | Transcription factor, SBP-box |
| SMART | SM00432 | 7.0E-39 | 889 | 948 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS50066 | 32.212 | 889 | 949 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 1.05E-36 | 890 | 947 | No hit | No description |
| SuperFamily | SSF55455 | 2.88E-28 | 890 | 951 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 891 | 945 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.7E-30 | 891 | 911 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 8.8E-25 | 899 | 945 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.7E-30 | 911 | 926 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.7E-30 | 926 | 947 | IPR002100 | Transcription factor, MADS-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0048653 | Biological Process | anther development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 974 aa Download sequence Send to blast |
MPLLKGEHGP GAPLGPSPLE SLQPSTSRKL CICCASSISL ASSLSLRALC SAQGTLVSLG 60 GLMDWDLKMP VSWDLAELEH DAVPAISAPT APVAAATSGI ATAAAAARGT PTRAECSVDL 120 KLGGLGDFGA AEVTKELKPP AKGPAVTVSS PPAAPAVVPS ASPLKRPRTG PGGANGPQCP 180 SCAVDGCDAD LSKCREYHRR HKVCEAHSKT PVVVVAGREM RFCQQCSRFH SLVEFDEAKR 240 SCRKRLDGHN RRRRKPQPDD MNARSFMTSQ QGPRLSSFVT PRPEPSWSGV MKLEDSSYSA 300 HQVLCNRPHF AGSTSTYSKE GRRFPFLQDG DQLSFGTGGA ASLEAATACQ SLLKTVAAPE 360 SSSSNKIFSP DRLTSVLDSD CALSLLSSSA NSSSVDVSRM VQPTEHIPMA QPLVPSMQQF 420 GSSPSWFSQA ASSGVVVSTA GFTCPSMDSE QLNPVLVPSS DGHEMNYHGI FHVGGEGSYD 480 GTSSSLPFSW HFDRRLLSSM AEFLFARPTL HSYHPCWIRN ASVFARSRRT AGRGWAVGWY 540 RLSARQDDSH ANLSAPIASR GRAGGRQGTK FCLARPGARR GRADKASEHG KMSVLGVQAS 600 KVEWHVTSGC AEQPPRATGR PPDLGIVDRH VVKLMTGSCR VRAPYDVAPH PAIPPPTARV 660 RARYPADAAG GPSNRWFRTA DDGVRVHNLG LAPLAEGWRE VSVLYSLVPL MTTEKPSRLT 720 LTGELRPVCS NCGNQSLASS RAEPERQPRR PVAGLEEAGN DAAHGKRKGG VAQIYDSTRG 780 RVDVRADHET GLGFRPLGFR VAHHLTRNGK ERSAPITPPP SRPAFTTPRR PAISTQRTSP 840 ALTSSVPHSS PPSLPLCVCC FAWREACLLL TSVGSRLAAT PAREEARAMG RGRVELKQIE 900 NKINRQVTFA KRRNGLLKKA YELSVLCDAE VALIIFSNRG KLFEFCSGQR YTAAYTHAAY 960 LRNEEMPVVG AADR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-27 | 170 | 255 | 1 | 84 | squamosa promoter binding protein-like 4 |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 238 | 255 | KRSCRKRLDGHNRRRRKP |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00555 | DAP | Transfer from AT5G50570 | Download |
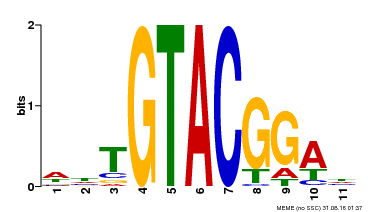
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc00017.1.g00700.1.am.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT053875 | 7e-93 | BT053875.1 Zea mays full-length cDNA clone ZM_BFb0379K14 mRNA, complete cds. | |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1755 | 37 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G50670.1 | 7e-44 | SBP family protein | ||||



