 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zpz_sc00003.1.g00180.1.am.mk | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | ERF | ||||||||
| Protein Properties | Length: 299aa MW: 33155.5 Da PI: 11.2165 | ||||||||
| Description | ERF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 52.3 | 1.4e-16 | 131 | 180 | 2 | 55 |
AP2 2 gykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
+y+GVr +k +g+WvAeIr+p + r+r++lg++ t+ Aa+a++ a +l+g
Zpz_sc00003.1.g00180.1.am.mk 131 QYRGVRMRK-WGKWVAEIREPHK---RTRIWLGSYATPVAAARAYDTAVFYLRG 180
69*****99.**********732...5*************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:3.30.730.10 | 3.4E-30 | 130 | 189 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.83E-21 | 131 | 190 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 7.9E-36 | 131 | 194 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 22.589 | 131 | 188 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 3.8E-10 | 131 | 180 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 8.8E-12 | 132 | 143 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.78E-24 | 134 | 189 | No hit | No description |
| PRINTS | PR00367 | 8.8E-12 | 170 | 190 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 299 aa Download sequence Send to blast |
MPPRTRKIPV QIPPSKWIAR ERREKARPTP KATASCCGSG LLPFRRPAYR YPSREGSRSR 60 SRRRRLGWLL MQGEYRSSSE ESAAAAAAAA AAAAAAMAPL AAAAAAAVKM EEQAAMAPSA 120 EQHQQQQPRR QYRGVRMRKW GKWVAEIREP HKRTRIWLGS YATPVAAARA YDTAVFYLRG 180 RSARLNFPDE IPALALSEGG DADGGTLSAA SIRKKAIEVG SRVDALQTGM MVAPPPHHCE 240 RPRHNHHHHH QHRELQLLGD EHHQHEQKPP PQQPRAAWSG RAKNPDLNRA PSPESSDAE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2gcc_A | 1e-13 | 130 | 188 | 4 | 63 | ATERF1 |
| 3gcc_A | 1e-13 | 130 | 188 | 4 | 63 | ATERF1 |
| 5wx9_A | 1e-13 | 131 | 190 | 14 | 74 | Ethylene-responsive transcription factor ERF096 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probably acts as a transcriptional activator. Binds to the GCC-box pathogenesis-related promoter element. May be involved in the regulation of gene expression by stress factors and by components of stress signal transduction pathways (By similarity). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00276 | DAP | Transfer from AT2G23340 | Download |
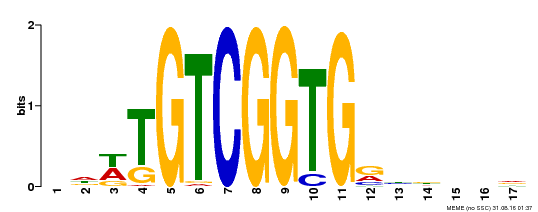
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Zpz_sc00003.1.g00180.1.am.mk |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT070130 | 1e-104 | BT070130.1 Zea mays full-length cDNA clone ZM_BFc0110K17 mRNA, complete cds. | |||
| GenBank | EU966906 | 1e-104 | EU966906.1 Zea mays clone 298090 dehydration responsive element binding protein mRNA, complete cds. | |||
| GenBank | KJ727878 | 1e-104 | KJ727878.1 Zea mays clone pUT5971 AP2-EREBP transcription factor (EREB47) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025826193.1 | 2e-80 | ethylene-responsive transcription factor RAP2-10-like | ||||
| Swissprot | O22174 | 5e-46 | ERF08_ARATH; Ethylene-responsive transcription factor ERF008 | ||||
| TrEMBL | A0A2S3IAZ3 | 3e-79 | A0A2S3IAZ3_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.J27349.1.p | 5e-84 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3943 | 30 | 75 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G23340.1 | 2e-41 | DREB and EAR motif protein 3 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



