 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc07475.1.g00010.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 778aa MW: 82330 Da PI: 7.9538 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 38.1 | 3.9e-12 | 383 | 439 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pseng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s+y+GV++++++gr++A+++d + ++ ++ k + g ++ +e+Aa+a++ a++k++g
Zmw_sc07475.1.g00010.1.am.mk 383 SQYRGVTRHRWTGRYEAHLWDnSC-RKegQTrKGRQ-GGYDMEEKAARAYDLAALKYWG 439
78*******************433.44555535555.77******************98 PP
| |||||||
| 2 | AP2 | 24.2 | 8.2e-08 | 482 | 516 | 1 | 38 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgt 38
s y+GV+++++ grW A+I +s +k +lg+f t
Zmw_sc07475.1.g00010.1.am.mk 482 SIYRGVTRHHQHGRWQARIGRVSG---NKDLYLGTFST 516
57****************999754...4********87 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| CDD | cd00018 | 1.53E-16 | 383 | 446 | No hit | No description |
| Pfam | PF00847 | 1.3E-9 | 383 | 439 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 1.18E-13 | 383 | 448 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 7.2E-21 | 384 | 453 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 16.937 | 384 | 447 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 1.2E-11 | 384 | 447 | IPR001471 | AP2/ERF domain |
| CDD | cd00018 | 2.68E-16 | 482 | 589 | No hit | No description |
| SuperFamily | SSF54171 | 7.19E-6 | 482 | 517 | IPR016177 | DNA-binding domain |
| SMART | SM00380 | 4.1E-16 | 483 | 593 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 13.485 | 483 | 587 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 7.8E-5 | 483 | 516 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.2E-6 | 561 | 587 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007276 | Biological Process | gamete generation | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0042127 | Biological Process | regulation of cell proliferation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 778 aa Download sequence Send to blast |
VLQDINSVVT VAREVKLKTG RSCARGDLYT TMRKTDCETT LLVDSAHAYL PPEVGSARLG 60 SGFLPTSHRH HLVLLRLACR RHYKMTSNSS QNVSSCSTGG GDAAAGGGSW LGFSLSPHHV 120 ASMDAAADGA RGVPQHHLGL FYPPVTSSPA PFCYALGGSG QDSVVGQAAG ASGGGFYPAL 180 SSMPLRSDGS LCITDAYRRS EEGQHHGMVA SSASPKLEDF LGAGPAMALS LDNSSLYYGG 240 GHGGHHGHGQ DHGGYPQPVH CAMIPGAAAT HDVYGHASMV DEQSAAAIAA SWFAARGVGG 300 YDVNSTGVDA GLVPCHPLAL SMSSGTGSQA SCVTMQMGAH ADPVAEYIAM EGSKKRGTDR 360 GAGAGQKQTV HRKSIDTFGQ RTSQYRGVTR HRWTGRYEAH LWDNSCRKEG QTRKGRQGGY 420 DMEEKAARAY DLAALKYWGP STHINFSLED YREELEEMKN MTRQEYVAHL RRKSSGFSRG 480 ASIYRGVTRH HQHGRWQARI GRVSGNKDLY LGTFSTCSVF HLLFLSIHPW PLPGECLLRR 540 FPHTFRRAFA SASPGASPSS PTQEEAAEAY DVAAIKFRGL NAVTNFDITR YDVEKIMESS 600 TLLPSELVRR RKDGDVGTVA AAAAITEGDA TAVANAAAAL VHAGNGSNAV EWKIQAAAAL 660 PAAPGRTDEH VQQNHGILAS EAFSAFSGLD DMVPGVEADN AHMSNASSLA TSVSNSREQS 720 PERGGGLAVL FAKPVAAASK LTCPPLPLGS WVSPSPVSAR PVVSIAHLPM FAAWTDAA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Acts as positive regulator of adventitious (crown) root formation by promoting its initiation. Promotes adventitious root initiation through repression of cytokinin signaling by positively regulating the two-component response regulator RR1. Regulated by the auxin response factor and transcriptional activator ARF23/ARF1. {ECO:0000269|PubMed:21481033}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00088 | SELEX | Transfer from AT4G37750 | Download |
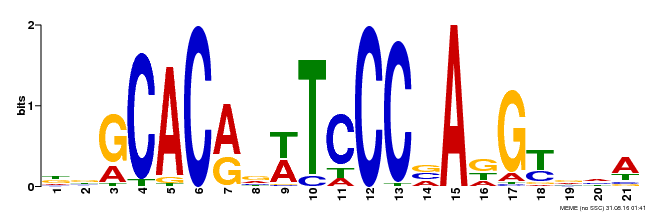
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by auxin. {ECO:0000269|PubMed:21481033}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025796942.1 | 0.0 | AP2-like ethylene-responsive transcription factor CRL5 | ||||
| Swissprot | Q84Z02 | 1e-155 | CRL5_ORYSJ; AP2-like ethylene-responsive transcription factor CRL5 | ||||
| TrEMBL | A0A1E5VRG8 | 0.0 | A0A1E5VRG8_9POAL; AP2-like ethylene-responsive transcription factor ANT | ||||
| STRING | Pavir.J24038.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP948 | 38 | 133 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



