 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc05640.1.g00010.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 375aa MW: 41799.5 Da PI: 6.9037 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 28.3 | 3e-09 | 300 | 338 | 17 | 55 |
HHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 17 elFeknrypsaeereeLAkklgLterqVkvWFqNrRake 55
e ++k +yps++++ LA+++gL+ +q+ +WF N+R ++
Zmw_sc05640.1.g00010.1.sm.mk 300 ETYYKWPYPSETQKVALAESTGLDLKQINNWFINQRKRH 338
578899******************************885 PP
| |||||||
| 2 | ELK | 32.5 | 1.9e-11 | 258 | 279 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELKh+Ll KYsgyL+sLkqE+s
Zmw_sc05640.1.g00010.1.sm.mk 258 ELKHHLLMKYSGYLSSLKQELS 279
9*******************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 8.3E-8 | 88 | 163 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 9.1E-11 | 89 | 112 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 1.6E-4 | 140 | 162 | IPR005540 | KNOX1 |
| SMART | SM01256 | 1.4E-25 | 170 | 221 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 2.8E-22 | 176 | 219 | IPR005541 | KNOX2 |
| SMART | SM01188 | 570 | 179 | 199 | IPR005539 | ELK domain |
| PROSITE profile | PS51213 | 10.807 | 258 | 278 | IPR005539 | ELK domain |
| SMART | SM01188 | 7.1E-7 | 258 | 279 | IPR005539 | ELK domain |
| Pfam | PF03789 | 7.5E-10 | 258 | 279 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.698 | 278 | 341 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 3.34E-19 | 279 | 354 | IPR009057 | Homeodomain-like |
| SMART | SM00389 | 2.1E-11 | 280 | 345 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 2.5E-26 | 283 | 344 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 3.14E-12 | 290 | 342 | No hit | No description |
| Pfam | PF05920 | 1.0E-14 | 298 | 337 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 316 | 339 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 375 aa Download sequence Send to blast |
MEEISHNHFG VGASAHGQHQ HNPWGSALSA VVAPPPPLTL NTAATGNSGA SSGGNNPVLQ 60 LANGGSLLDA CVKAKEPSSS SPYAGDVEAI KAKIISHPHY YSLLAAYLEC QKASPTLLLL 120 LLHEKLLLHL MSNMEHAACN CKLVGAPPEV SARLTAMAQE LEARQRTSLL GLGSATEPEL 180 DQFMEAYHEM LVKFREELTR PLQEAMEFMR RVESQLGSLS ISSRSLRNIL SSGSEEDQEC 240 SGGETQLPEV DAHGVDQELK HHLLMKYSGY LSSLKQELSK KKKKGKLPKE ARQKLLSWWE 300 TYYKWPYPSE TQKVALAEST GLDLKQINNW FINQRKRHWK PSEEMHYMMM DGYHTSNAFY 360 MDGHFFNEAG MYRFG |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to RNA (PubMed:17965274). Possible transcription factor that regulates genes involved in development. Mutations in KN-1 alter leaf development. Foci of cells along the lateral vein do not differentiate properly but continue to divide, forming knots. May participate in the switch from indeterminate to determinate cell fates. Probably binds to the DNA sequence 5'-TGAC-3'. {ECO:0000269|PubMed:17965274}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | Transfer from GRMZM2G017087 | Download |
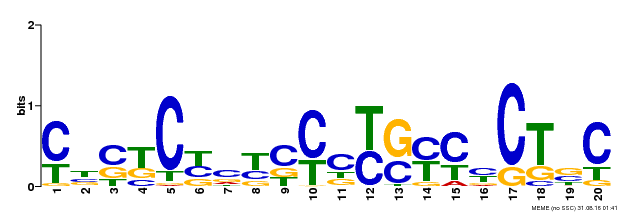
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004981913.1 | 0.0 | homeotic protein knotted-1 | ||||
| Swissprot | P24345 | 0.0 | KN1_MAIZE; Homeotic protein knotted-1 | ||||
| TrEMBL | A0A368SF91 | 0.0 | A0A368SF91_SETIT; Uncharacterized protein | ||||
| STRING | Pavir.J00864.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP9886 | 31 | 45 |



