 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc05542.1.g00030.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 847aa MW: 92455 Da PI: 5.8697 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.9 | 1.7e-18 | 14 | 72 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++++++ps +r++L + + +++ +q+kvWFqNrR +ek+
Zmw_sc05542.1.g00030.1.sm.mkhc 14 KYVRYTPEQVEALERLYYECPKPSSLRRQQLVRDCpvlaNVDPKQIKVWFQNRRCREKQ 72
6789*****************************************************97 PP
| |||||||
| 2 | START | 173.1 | 1.8e-54 | 167 | 374 | 2 | 204 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakae 80
+ae++++e+++ka+ ++ Wv+++ +++g++++ +++ s+++ g a+ra+g+v m++a v+e+l+d++ W ++++++e
Zmw_sc05542.1.g00030.1.sm.mkhc 167 IAEQTLTEFLSKATGTAVEWVQMPGMKPGPDSIGIIAISHGCAGVAARACGLVGMEPA-KVAEILKDRPLWLRDCRSME 244
689999****************************************************.8888888888********** PP
EEEEECTT..EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEE CS
START 81 tlevissg..galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgil 153
+++v+ g g+++l +++l+a+++l+p Rdf+ +Ry+ l +g++v++++S++s+q p+ ++++R+e+lpSg+l
Zmw_sc05542.1.g00030.1.sm.mkhc 245 VVNVLPAGnsGTIELLYMQLYAPTTLAPaRDFWLLRYTSILDDGSLVVCERSLSSKQGGPTmplVQPFIRGEMLPSGFL 323
************************************************************999999************* PP
EEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHHHHHHHHHHHHHHTXXXX CS
START 154 iepksnghskvtwvehvdlkgrlphwllrslvksglaegaktwvatlqrqc 204
i+p+++g+s +++v+h+dl+ +++++++r+l++s+++ ++k+ +a+l++++
Zmw_sc05542.1.g00030.1.sm.mkhc 324 IRPSDGGGSVIHIVDHMDLEPWSVPEVVRPLYESSAMVAQKMSMAALRYLR 374
***********************************************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.337 | 9 | 73 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-14 | 11 | 77 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 4.0E-16 | 14 | 72 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.01E-15 | 14 | 74 | No hit | No description |
| SuperFamily | SSF46689 | 2.44E-16 | 15 | 75 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 1.3E-17 | 16 | 72 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 4.14E-6 | 66 | 105 | No hit | No description |
| PROSITE profile | PS50848 | 25.951 | 157 | 376 | IPR002913 | START domain |
| CDD | cd08875 | 2.96E-73 | 161 | 375 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 5.2E-22 | 164 | 371 | IPR023393 | START-like domain |
| SMART | SM00234 | 6.2E-45 | 166 | 376 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 2.75E-36 | 166 | 376 | No hit | No description |
| Pfam | PF01852 | 3.6E-52 | 167 | 374 | IPR002913 | START domain |
| Pfam | PF08670 | 2.3E-49 | 703 | 846 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 847 aa Download sequence Send to blast |
MVTAKEAAAM DASKYVRYTP EQVEALERLY YECPKPSSLR RQQLVRDCPV LANVDPKQIK 60 VWFQNRRCRE KQRKESSRLQ ALNRKLTAMN KLLMEENDRL QKQVSQLVYE NGYYRQQTQS 120 AGLATTDTSC ESVVTSGQQN VAAAAAAAAA APQAQPRDAS PAGLMSIAEQ TLTEFLSKAT 180 GTAVEWVQMP GMKPGPDSIG IIAISHGCAG VAARACGLVG MEPAKVAEIL KDRPLWLRDC 240 RSMEVVNVLP AGNSGTIELL YMQLYAPTTL APARDFWLLR YTSILDDGSL VVCERSLSSK 300 QGGPTMPLVQ PFIRGEMLPS GFLIRPSDGG GSVIHIVDHM DLEPWSVPEV VRPLYESSAM 360 VAQKMSMAAL RYLRQVANED TQSVITGWGR QPTALRALSQ KLTRCYNEAI NGLADDGWSV 420 IESDGVDDVC ISVNSSPTKV INCNATFNNG LPIVSSSVLC AKASMLLQDV SPASLLRFMR 480 EQRSQWADSS LDAFFASAMK PNFCNLPMSR LGGFSGQVIL PLAHMFDPEE FLEIIKLGNA 540 SNYQDALMHR DLFLLQMYNG VDENTLGTCS ELIFAPIDAS FSDDSPLLPS GFRIIPIDSP 600 LDSSSPNCTL DLASTLEVGT PRSRVTGNGP ANSSCASSKS VMTIAFQFAF ERHLQDSVAT 660 MARQYVRSII QSVQRIALAL SSSRPIPQCG VSHTPAISPE AATLSIWICQ SYRFHFGAEL 720 IKSADSGSCE PSLKALWHHA SAVLCCSLKA MPVFSFANQS GLDMLETTLV SLQDITLEKV 780 FDDQGRKNLC TELPTIMEQG FACIPGGLCV SSLGRPVSYE KALAWKVLDD VNSAHCICFM 840 FVNWSFV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
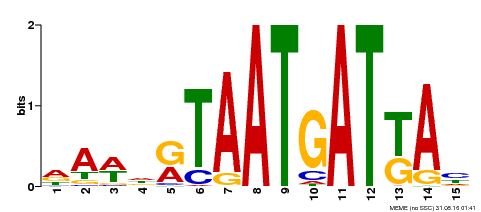
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025816436.1 | 0.0 | homeobox-leucine zipper protein HOX29-like | ||||
| Swissprot | A2WLR5 | 0.0 | HOX29_ORYSI; Homeobox-leucine zipper protein HOX29 | ||||
| TrEMBL | A0A3L6QYP2 | 0.0 | A0A3L6QYP2_PANMI; Homeobox-leucine zipper protein HOX29-like isoform X2 | ||||
| STRING | Pavir.Eb00113.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP374 | 38 | 197 |



