 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc04648.1.g00070.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 275aa MW: 30355.6 Da PI: 9.2445 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 107.1 | 9.3e-34 | 26 | 80 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kprlrWtp+LHerFveav++LGG++kAtPk++l+lm++kgLtl+h+kSHLQkYRl
Zmw_sc04648.1.g00070.1.sm.mkhc 26 KPRLRWTPDLHERFVEAVTKLGGPDKATPKSVLRLMGMKGLTLYHLKSHLQKYRL 80
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.032 | 23 | 83 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.0E-33 | 23 | 81 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 9.32E-17 | 26 | 83 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 8.6E-25 | 26 | 81 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 8.4E-10 | 28 | 79 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 3.7E-24 | 125 | 169 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 275 aa Download sequence Send to blast |
MFEGMERAGY GAAAAMGGVV LSRDPKPRLR WTPDLHERFV EAVTKLGGPD KATPKSVLRL 60 MGMKGLTLYH LKSHLQKYRL GKQNKKDTGL EARRGAFAAQ GINFSAPAPP TIASAASNST 120 GETPLADALK YQIEVQRKLH EQLEVQKKLQ MRIEAQGKYL QTILEKAQSN LSYDASGAEN 180 LEATRSQLTD FNLALSGFMD NVTQVCEQNS GELTKAMSQD NLRTSNIGFQ LYHGVQDVED 240 VKCTPDDGLL LLDLNIRGGY GHRPLSDLKI NQHMR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 4e-21 | 26 | 82 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 4e-21 | 26 | 82 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 4e-21 | 26 | 82 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 4e-21 | 26 | 82 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
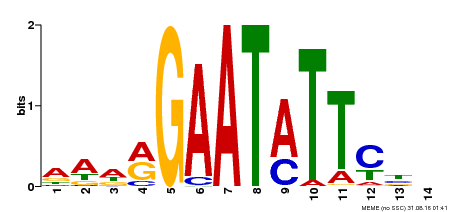
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00544 | DAP | Transfer from AT5G45580 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025808925.1 | 1e-177 | myb family transcription factor PHL11-like | ||||
| TrEMBL | A0A2S3GSU5 | 1e-176 | A0A2S3GSU5_9POAL; Uncharacterized protein | ||||
| STRING | Si018049m | 1e-176 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3940 | 34 | 70 |



