 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc02538.1.g00140.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 263aa MW: 29000.8 Da PI: 8.8339 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 40.9 | 3.6e-13 | 83 | 128 | 4 | 54 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksL 54
+h e+Er+RR+++N++f Lr ++P+ K++Ka+ L A+ YI++L
Zmw_sc02538.1.g00140.1.es.mkhc 83 NHVEAERQRREKLNQRFYALRAVVPNV-----SKMDKASLLGDAISYINEL 128
799***********************6.....5****************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 17.39 | 79 | 128 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 9.81E-19 | 82 | 153 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.25E-12 | 82 | 132 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 6.4E-19 | 83 | 150 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 1.2E-10 | 83 | 128 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 5.6E-16 | 85 | 134 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006952 | Biological Process | defense response | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043425 | Molecular Function | bHLH transcription factor binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 263 aa Download sequence Send to blast |
MEATSRASNT NLHPAATANE GMLSFSSTPT TRPAKSESDH SDLDASVREV ESSRVVEPPP 60 PEAEKRPRKR GRKPANGREE PLNHVEAERQ RREKLNQRFY ALRAVVPNVS KMDKASLLGD 120 AISYINELRG KLTALESDKE TMQSQIEALK KERDARPAPQ AGGFGHDAGP RCHAVEIDAK 180 ILGLEAMIRV HCHKRNHPSA RLMTALRELD LDVCHASVSV VEDLMIQQVA VKMASRIYSQ 240 NQLNAALYSR LAEPGSVVGR ESS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 5e-35 | 77 | 156 | 2 | 81 | Transcription factor MYC2 |
| 5gnj_B | 5e-35 | 77 | 156 | 2 | 81 | Transcription factor MYC2 |
| 5gnj_E | 5e-35 | 77 | 156 | 2 | 81 | Transcription factor MYC2 |
| 5gnj_F | 5e-35 | 77 | 156 | 2 | 81 | Transcription factor MYC2 |
| 5gnj_G | 5e-35 | 77 | 156 | 2 | 81 | Transcription factor MYC2 |
| 5gnj_I | 5e-35 | 77 | 156 | 2 | 81 | Transcription factor MYC2 |
| 5gnj_M | 5e-35 | 77 | 156 | 2 | 81 | Transcription factor MYC2 |
| 5gnj_N | 5e-35 | 77 | 156 | 2 | 81 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 64 | 72 | KRPRKRGRK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator involved in jasmonate (JA) signaling pathway during spikelet development. Binds to the G2 region G-box (5'-CACGTG-3') of the MADS1 promoter and thus directly regulates the expression of MADS1. Its function in MADS1 activation is abolished by TIFY3/JAZ1 which directly target MYC2 during spikelet development. {ECO:0000269|PubMed:24647160}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00085 | PBM | Transfer from AT5G46760 | Download |
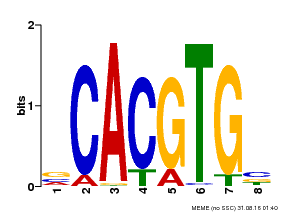
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_012698381.1 | 1e-164 | LOW QUALITY PROTEIN: transcription factor MYC2-like | ||||
| Swissprot | Q336P5 | 1e-152 | MYC2_ORYSJ; Transcription factor MYC2 | ||||
| TrEMBL | A0A1E5W733 | 1e-164 | A0A1E5W733_9POAL; Transcription factor MYC2 | ||||
| STRING | Pavir.J38781.1.p | 1e-167 | (Panicum virgatum) | ||||
| STRING | Si039973m | 1e-164 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4452 | 31 | 57 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



