 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc01754.1.g00080.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 299aa MW: 32228.5 Da PI: 7.2155 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 40.4 | 6.6e-13 | 75 | 119 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r +WT+ E++++++a +++ + Wk+I +++g +t q++s+ qky
Zmw_sc01754.1.g00080.1.sm.mkhc 75 RESWTEPEHDKFLEALQLFDRD-WKKIEAYVG-SKTVIQIRSHAQKY 119
789*****************77.*********.*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.17E-16 | 69 | 125 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 14.37 | 70 | 124 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 1.0E-18 | 73 | 122 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 6.8E-6 | 74 | 115 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 2.7E-10 | 74 | 122 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.1E-10 | 75 | 119 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 6.60E-8 | 77 | 120 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 299 aa Download sequence Send to blast |
MVSMSTTPMP LDAVKSAAVA GGGVDEGGGG GGKQQLMRGA AAAIAPPPMA VPAPVPAAGE 60 EVRKVRKPYT ITKSRESWTE PEHDKFLEAL QLFDRDWKKI EAYVGSKTVI QIRSHAQKYF 120 LKVEKNGTGE HLPPPRPKRK AAHPYPQKAS KTVSQAVLSQ QLHPLRDQGS VMSMDTSNVV 180 RNTNPNGAGT YCDNALVQPF SASQGVVVTN NCSSSIEGPS GNWPASEAVE QENVVPALRA 240 MPDFARVYSF LGSIFDPDTS GHLQRLKAMD PIDVETVLLL MRNLSTNLTS PDFEEHGNE |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00560 | DAP | Transfer from AT5G52660 | Download |
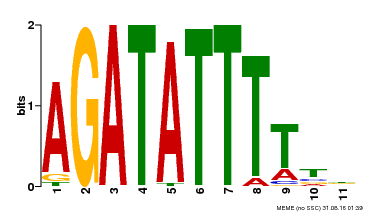
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025812931.1 | 1e-156 | protein REVEILLE 6-like | ||||
| Swissprot | Q8H0W3 | 8e-90 | RVE6_ARATH; Protein REVEILLE 6 | ||||
| TrEMBL | A0A2T7DVG3 | 1e-156 | A0A2T7DVG3_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Da00204.1.p | 1e-155 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1096 | 38 | 111 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



