 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc01419.1.g00200.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 666aa MW: 70575.4 Da PI: 9.4085 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 49.8 | 6.1e-16 | 471 | 516 | 5 | 55 |
HHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 5 hnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hne Er+RRdriN+++ +L++l+P++ K +Ka++L ++++Y+k+Lq
Zmw_sc01419.1.g00200.1.am.mk 471 HNESERKRRDRINQKMKTLQKLVPNS-----SKTDKASMLDEVIDYLKQLQ 516
*************************8.....7******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50888 | 18.025 | 466 | 515 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 1.24E-19 | 469 | 539 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene3D | G3DSA:4.10.280.10 | 1.3E-17 | 471 | 528 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.48E-17 | 471 | 520 | No hit | No description |
| Pfam | PF00010 | 1.7E-13 | 471 | 516 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.2E-18 | 472 | 521 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009567 | Biological Process | double fertilization forming a zygote and endosperm | ||||
| GO:0009506 | Cellular Component | plasmodesma | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 666 aa Download sequence Send to blast |
MAPAKFGSST RSRRNKSKTG SNTRDEQASR LRATTRWLSH RSYRRQPARH ALNLPYAGSS 60 DWERWPRGHV EAEAATCPPP PHKAQAPRGL VFLRAVTLTA PASTYPHARS RPSPIDGARC 120 MAPATHHPAY PPPPVLKYLL LRAIRSPRPP HRASTRALAA RSNRALPASL TAAPVSSRLT 180 SSHATGRGGG AAEMNQCVPS WDLDDAAAGG LNTASAATDG VHHRAGVDGA GYYDVAELTW 240 EKGNISSHGL LNRPAPTKCP PAASSTPLQA LGGGGEPAGG GDRETLEAVV GEAAARSHFP 300 AQPSLPTTPW LGVGVPLPAE ALVPCATRDD QAAEEEARRK RARGVGEDGH GLVCASQGST 360 APGRRGESAL PELDAACGIG DDDVCGFTTT TNNSTSLDRD DNKGSPDTEN TSIGGGASDS 420 RCFSLRSQAN CLLITLPRDG LCDEGENVVI NGEPAMRSSV STKRSRAAAI HNESERKRRD 480 RINQKMKTLQ KLVPNSSKTD KASMLDEVID YLKQLQAQVQ VMSRMSSMMM PMAMPQLQMS 540 VMAQMAQMAQ MAQMAQGMMN MGSLSQPGYV PPPMMHPPPP PFVPLPWDAG AAGASSGAGA 600 VADRVPQPGG SATITDAFAA FLACQQAQQN GQQQQQPGGM EAYNRMLAMW HKLSQQQPSQ 660 PSNTKP |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 474 | 479 | ERKRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00034 | PBM | Transfer from AT4G00050 | Download |
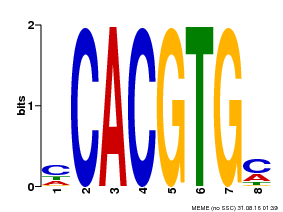
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_014660002.1 | 1e-151 | transcription factor UNE10 isoform X2 | ||||
| TrEMBL | A0A368SN18 | 1e-149 | A0A368SN18_SETIT; Uncharacterized protein | ||||
| STRING | Pavir.Ia02188.1.p | 1e-149 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7474 | 31 | 40 |



