 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc01402.1.g00270.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 440aa MW: 47179.9 Da PI: 8.7863 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 98.3 | 5.4e-31 | 265 | 320 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
kpr++W+peLH+rF++a+ qLGGs++AtPk+i++lmkv+gLt+++vkSHLQkYRl+
Zmw_sc01402.1.g00270.1.sm.mk 265 KPRRCWAPELHRRFLQALLQLGGSHVATPKQIRDLMKVDGLTNDEVKSHLQKYRLH 320
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 12.466 | 262 | 322 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-27 | 262 | 323 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.49E-16 | 263 | 323 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.6E-25 | 265 | 320 | IPR006447 | Myb domain, plants |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 440 aa Download sequence Send to blast |
MEMDPPPPSL HAAAAAGDRR RCREYLLALE EERRKIQVFQ RELPLCLDLV TQTIERMKSQ 60 MDGIGSEVTV SDHGPVLEEF MPLKPIRSLS SSEEHDSARD ASGGVGRKEE AADTPAETKK 120 ATPDWLQSVP LWSQEPQQRS ASDPHKVHTD MFKPCSLPVF RLQIITPLTI DTSQLYQELP 180 CKPVALNARK SGGAFQPFEK EKRVELPASS TTAAASSTVA GDSGDDDKGA SVDTEKRSGE 240 KETSTNKDAK GAKDKEGQSS SSSRKPRRCW APELHRRFLQ ALLQLGGSHV ATPKQIRDLM 300 KVDGLTNDEV KSHLQKYRLH TRRPNSTGQS SSASAAPPAP QFVVVGGIWV PPPEYAAAAA 360 AVAAAAAAQP PVQQLAANTS GGASTVYAPV ATLPSAPQPR SQQLVKKQPN RCSDGRRRGS 420 AGETSSASPA VSSSSQTTSA |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor that may play a role in response to nitrogen. May be involved in a time-dependent signaling for transcriptional regulation of nitrate-responsive genes. Binds specifically to the DNA sequence motif 5'-GAATC-3' or 5'-GAATATTC-3'. Represses the activity of its own promoter trough binding to these motifs. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
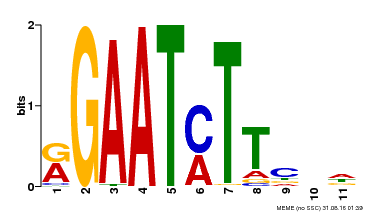
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by nitrate. {ECO:0000269|PubMed:23324170}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_004954868.3 | 1e-159 | transcription factor NIGT1 | ||||
| Swissprot | Q6Z869 | 1e-132 | NIGT1_ORYSJ; Transcription factor NIGT1 | ||||
| TrEMBL | A0A0A9G3F3 | 1e-175 | A0A0A9G3F3_ARUDO; Uncharacterized protein | ||||
| STRING | Si019945m | 1e-160 | (Setaria italica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6470 | 37 | 54 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



