 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc00911.1.g00130.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 492aa MW: 54222.8 Da PI: 7.6442 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 53.8 | 3.3e-17 | 229 | 282 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rqV +WFqNrRa++k
Zmw_sc00911.1.g00130.1.am.mk 229 KKRRLNVEQVRTLEKNFELRNKLEPERKLQLARALGLQPRQVAIWFQNRRARWK 282
4568999**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 125.5 | 2.4e-40 | 228 | 319 | 1 | 92 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLe 81
ekkrrl+ eqv++LE++Fe ++kLeperK +lar+Lglqprqva+WFqnrRAR+ktkqlEkdy+aLkr++da++++n++L
Zmw_sc00911.1.g00130.1.am.mk 228 EKKRRLNVEQVRTLEKNFELRNKLEPERKLQLARALGLQPRQVAIWFQNRRARWKTKQLEKDYDALKRQLDAVQADNDALL 308
69******************************************************************************* PP
HD-ZIP_I/II 82 keveeLreelk 92
+++++L++e++
Zmw_sc00911.1.g00130.1.am.mk 309 SQNKKLQAEVM 319
*******9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.3E-18 | 211 | 285 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.56E-19 | 220 | 286 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.443 | 224 | 284 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.2E-16 | 227 | 288 | IPR001356 | Homeobox domain |
| Pfam | PF00046 | 1.3E-14 | 229 | 282 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 1.15E-15 | 229 | 285 | No hit | No description |
| PRINTS | PR00031 | 3.1E-5 | 255 | 264 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 259 | 282 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 3.1E-5 | 264 | 280 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.4E-11 | 284 | 320 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 492 aa Download sequence Send to blast |
MEKESKWKRE REDERVQVEA ARSWENEQGH VRLWPRTNAG AGQARRGAPE VVGPHPRAVY 60 AALPLLQQLR SLVVNERGRG HEGANASHNL ASSVIHLRLQ RGRVHHTFTP VTYAHLRPMA 120 SNGMASSPSF FPPNFLLQMQ QAPPDHEPQE HHQHHHHLPP PPLHPHHNPF LPSSQCPSLQ 180 DFRGMAPMLG KRPMYGADVG GGDEVNGCGG GGGANEDELS DDGSQAGEKK RRLNVEQVRT 240 LEKNFELRNK LEPERKLQLA RALGLQPRQV AIWFQNRRAR WKTKQLEKDY DALKRQLDAV 300 QADNDALLSQ NKKLQAEVMS RLTLAELGSM WVSVLAAKRL LEAAAIGAAF SPARVILELK 360 GGREAGSELI NLNKKTEASC SNRSENSSEI NLDISRTPPS EGAMDPPAHH HPEHASNGGG 420 MIPFYPSSGR IAGVDIDQLL HSSSGPKIEQ QGNGARAGVQ APETASFSNL LCGVDEPPPF 480 WPWADHQPSI DD |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 276 | 284 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
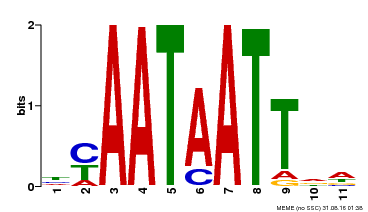
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002465782.1 | 1e-173 | homeobox-leucine zipper protein HOX21 | ||||
| Swissprot | Q8S7W9 | 1e-126 | HOX21_ORYSJ; Homeobox-leucine zipper protein HOX21 | ||||
| TrEMBL | A0A3L6SDB5 | 1e-178 | A0A3L6SDB5_PANMI; Uncharacterized protein | ||||
| STRING | Pavir.Ia04323.1.p | 1e-180 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2289 | 38 | 94 |



