 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zmw_sc00909.1.g00270.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Zoysieae; Zoysiinae; Zoysia
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 716aa MW: 78012.2 Da PI: 6.6772 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 49.1 | 1.3e-15 | 24 | 68 | 1 | 47 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
r rWT+ E++++++a k++G W +I +++g ++t+ q++s+ qk+
Zmw_sc00909.1.g00270.1.am.mkhc 24 RERWTEAEHKRFLEALKLYGRQ-WQRIEEHVG-TKTAVQIRSHAQKF 68
78******************77.*********.************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 2.47E-16 | 18 | 74 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 19.824 | 19 | 73 | IPR017930 | Myb domain |
| TIGRFAMs | TIGR01557 | 2.2E-16 | 22 | 71 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.6E-12 | 23 | 71 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.1E-8 | 24 | 64 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 5.8E-13 | 24 | 67 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 2.38E-9 | 26 | 69 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 716 aa Download sequence Send to blast |
MEVNSSGEET VVKVRKPYTI TKQRERWTEA EHKRFLEALK LYGRQWQRIE EHVGTKTAVQ 60 IRSHAQKFFT KLEKEAMNNG TSPGKAHDID IPPPRPKRKP NSPYPRKNGL NSETSTKEVS 120 NDKSTKSNIR LSSGNVEMAS HSSLQKLQRK EASEKGSCSE VLNLFREVPS ASFCSVNKTY 180 SNHGAPSGIE QAKTERDMTT LGKNPLSIDA HEGAKEINGS EMEKANGILI SSKSDRPHED 240 YLDTSTQEMK LKPMSVDTAY VGQQTAKTSH SVTEKNGATS VLATGTDGSH ADQTSDHDGA 300 NGRMNPCIHP TFSVDPKFNI NAAPLPCPHN YTALPPMMQC NCNQDAYRSF INMSSTLSSM 360 LVSTLLSNPA IHAAARLAAS YWPAAENNTS VDPNQENPTE CVQGSNIGSP QSMASMVAAT 420 VAAASAWWAT QGLLPFFTPP MAFPFAPAPS TAFPTADVRA SEKGRDCQVE NAQKECQEAQ 480 NQDQSESLRV AASSESDRSG KGDLSLQTEL KISPVQIADT TPTTVADTID AFKKKQDRSS 540 CGSNTPSSSD VEGDNVPGKQ DKGNDNAKQA SCSNSSAGDT NNRRFRSSGS TSDSWKEVSE 600 EGRLAFDALF SREKLPQSFS PPQGGESKEI SKEEEDEVTT VTVDLNKNAI AFDHDLDTMD 660 KSRASFSNEL SQLKLKSRRT GFKPYKRCSV EAKDNRVPAS DDVGTKRIRL ESEAST |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00654 | PBM | Transfer from LOC_Os08g06110 | Download |
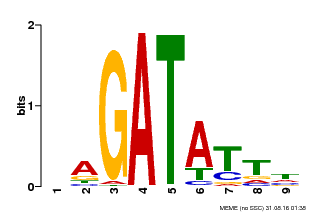
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025820794.1 | 0.0 | protein LHY isoform X1 | ||||
| Refseq | XP_025820795.1 | 0.0 | protein LHY isoform X1 | ||||
| TrEMBL | A0A2T8IF63 | 0.0 | A0A2T8IF63_9POAL; Uncharacterized protein | ||||
| STRING | Pavir.Fa01905.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6031 | 34 | 47 |



