 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma88g00640.1 | ||||||||
| Common Name | ZOSMA_88G00640 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 272aa MW: 30821.5 Da PI: 4.9576 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 43.3 | 8.7e-14 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd +l ++++G +W++ +r g+ R++k+c++rw +yl
Zosma88g00640.1 14 KGSWTAEEDMILMLHIQRYGHANWRSLPRQAGLFRCGKSCRLRWINYL 61
799*******************************************97 PP
| |||||||
| 2 | Myb_DNA-binding | 55.9 | 9.6e-18 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T eE+e ++++++++G++ W++Ia++++ gRt++++k+ w++yl
Zosma88g00640.1 67 RGNFTVEEEETIIKLHQKFGNR-WSAIAAELP-GRTDNEIKNVWHTYL 112
89********************.*********.*************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 3.0E-23 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 12.625 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 8.07E-30 | 11 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 6.9E-11 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 4.2E-12 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.29E-8 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 26.127 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.4E-27 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.1E-16 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.4E-16 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 8.22E-12 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:2000652 | Biological Process | regulation of secondary cell wall biogenesis | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 272 aa Download sequence Send to blast |
MVRAPCCEKI GLKKGSWTAE EDMILMLHIQ RYGHANWRSL PRQAGLFRCG KSCRLRWINY 60 LRPDLKRGNF TVEEEETIIK LHQKFGNRWS AIAAELPGRT DNEIKNVWHT YLKKRLMNDD 120 KKPAVGTMGR KSRNKNDHIP VDFQTNLIGS TSTSTSSGGS IFDQKQQPSQ SSDVSSDSSA 180 RGYVNVLEED EMMFMELPEI DDNFLLPNLP MDQLDTIVAD FPDYDDIVPV LSNGTTSDGE 240 EKDNHQNTED YMDFWLRVFL ESGSTGTDSS A* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 8e-26 | 12 | 116 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00375 | DAP | Transfer from AT3G23250 | Download |
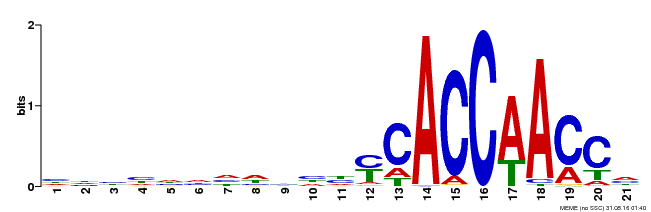
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010920477.1 | 5e-80 | transcription factor MYB30 | ||||
| TrEMBL | A0A0K9NKE8 | 0.0 | A0A0K9NKE8_ZOSMR; Myb domain protein 15 | ||||
| STRING | XP_008791593.1 | 1e-76 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP79 | 38 | 563 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G31180.1 | 6e-67 | myb domain protein 14 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma88g00640.1 |



