 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma5g00950.1 | ||||||||
| Common Name | ZOSMA_5G00950 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 986aa MW: 109715 Da PI: 6.5079 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 129.2 | 1.5e-40 | 140 | 217 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+CqvegC adl ak+yhrrhkvCe+h+ka++++v + qrfCqqCsrfh l+efDe+krsCrrrL++hn+rrrk+++
Zosma5g00950.1 140 MCQVEGCGADLIGAKDYHRRHKVCETHAKASKAIVGNAVQRFCQQCSRFHLLQEFDEGKRSCRRRLDGHNKRRRKAHP 217
6**************************************************************************875 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 8.8E-33 | 135 | 202 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.624 | 138 | 215 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.83E-36 | 140 | 219 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 9.4E-30 | 141 | 214 | IPR004333 | Transcription factor, SBP-box |
| CDD | cd00204 | 9.80E-8 | 758 | 884 | No hit | No description |
| SuperFamily | SSF48403 | 9.12E-7 | 787 | 885 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50297 | 8.747 | 788 | 846 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 4.4E-6 | 788 | 884 | IPR020683 | Ankyrin repeat-containing domain |
| PROSITE profile | PS50088 | 8.656 | 825 | 846 | IPR002110 | Ankyrin repeat |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 986 aa Download sequence Send to blast |
MDSQRLNGNG KKVMEWDLND WNWNGDLFLA TPVNASPPGC GNKRSFPSIL TTEVGLSTST 60 PSCSDETVHH GVNDKERYEL EEKRRKVIFV DDDEPVLEVS SLALKLGGIV HPKENEVING 120 NRIDNKSGKI QTPGGSSRGM CQVEGCGADL IGAKDYHRRH KVCETHAKAS KAIVGNAVQR 180 FCQQCSRFHL LQEFDEGKRS CRRRLDGHNK RRRKAHPDVV CTVPENPLTD AWSSSYLLIS 240 LLRILSNLQS KNTEPLNDQD LLTHFLKNLA SLAGSSDGKN LSSLMQACQD PLKVANSTGF 300 SSVVAEAEAA AAAPSNDVAL QESFRPFCFN SRSNPVTEVE EVNNLHKTST NESINPALIT 360 ELTGKRSIAK FFPESTINGV QKTQMVKLKS FDLNDAYNVE EEDIEIPEEV VAPRASLLNS 420 HLYMIQESHQ STSPPQTSGN SDSTSVQSFV SSNGDAKGRT DKIIFKLFGK DPKDLSPFIR 480 AQMFNWLSNT PSEIESYIKP GCIILTVYLC LSESYWLELC HNLSSSLKML LDLDSNGFWR 540 TGWMYVRVRN QMAFIYNGQL LLNSPYLGVS DHFTISRISP IAVSPSKTIT FRVNGFGSSL 600 SSARLFCAFE GKYLAQETTS SLVSSNGDVN SLNNGHGYLS FSCSIPDAVG RGFIEIEDES 660 LSSSFFPYII AEKEICSEIC MLEDTLEFDT HTDSTDISSE KLNNRSEALE FLHEMGWLLR 720 RSQLRSRTEE ICPGSDTFSI KRFKCVIEFG MKHEWRAVVK KLLDLLFGGV VDLSGCSIQL 780 VLGELALLHF AVQINSQPIV DLLLNYSKNE HKGILFRPDM VAIDTGLTPL HMAASRDDTE 840 AVLDALTDDP GQVGISAWKT IRDITNYTPE DYARAKNHES YLRLVDKKIT QKSNSSSTHV 900 IVDIPMSVFD IKKGNTPYCQ TCNQQQLKYM RKIQRRSLLI YKPAILSMVG IAAVCVCLGL 960 LLKGLPEVQY VLSPFRWETT RFGPI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-32 | 132 | 214 | 2 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
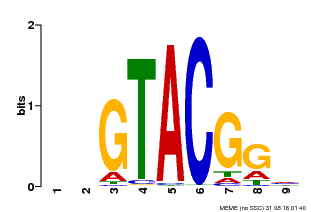
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010941481.1 | 0.0 | squamosa promoter-binding-like protein 6 | ||||
| TrEMBL | A0A0K9NW96 | 0.0 | A0A0K9NW96_ZOSMR; Putative Squamosa promoter-binding protein | ||||
| STRING | XP_008813721.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1998 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G60030.1 | 0.0 | squamosa promoter-binding protein-like 12 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma5g00950.1 |



