 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma4g02000.1 | ||||||||
| Common Name | ZOSMA_4G02000 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 520aa MW: 59173.7 Da PI: 6.9905 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 81.1 | 1.3e-25 | 205 | 258 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+r+ Wt +LH++Fv+av+q+ G +k Pk+il+lm+v+g+t+e+v+SHLQkYRl
Zosma4g02000.1 205 KARVIWTVDLHQKFVDAVNQI-GFDKVGPKKILDLMNVPGITRENVASHLQKYRL 258
68*******************.9*******************************8 PP
| |||||||
| 2 | Response_reg | 81.6 | 2.5e-27 | 22 | 130 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTT CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealkaG 93
vl+vdD+p+ +++l+++l+k y ev+++ +++ale+l++k+ +D+++ D++mp+mdG++ll++ e +lp+i+++ ge + + + ++ G
Zosma4g02000.1 22 VLVVDDDPTWLKILEKMLRKCSY-EVTTCGLARAALEMLRTKRgvFDIVISDVNMPDMDGFKLLEHVGLEM-DLPVIMMSVDGETSRVMKGVQHG 114
89*********************.******************99***********************6644.8********************** PP
ESEEEESS--HHHHHH CS
Response_reg 94 akdflsKpfdpeelvk 109
a d+l Kp+ ++el++
Zosma4g02000.1 115 ACDYLLKPVRMKELRN 130
*************987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 1.9E-142 | 1 | 517 | IPR017053 | Response regulator B-type, plant |
| Gene3D | G3DSA:3.40.50.2300 | 5.5E-45 | 19 | 162 | No hit | No description |
| SuperFamily | SSF52172 | 8.11E-36 | 19 | 157 | IPR011006 | CheY-like superfamily |
| SMART | SM00448 | 2.3E-32 | 20 | 132 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 43.125 | 21 | 136 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 3.5E-24 | 22 | 131 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 4.25E-30 | 23 | 136 | No hit | No description |
| SuperFamily | SSF46689 | 1.61E-18 | 203 | 262 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.6E-28 | 204 | 263 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.0E-22 | 205 | 258 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 4.5E-7 | 210 | 257 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0000160 | Biological Process | phosphorelay signal transduction system | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009735 | Biological Process | response to cytokinin | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 520 aa Download sequence Send to blast |
MMEAPSSCSS SRAEAFPAGL RVLVVDDDPT WLKILEKMLR KCSYEVTTCG LARAALEMLR 60 TKRGVFDIVI SDVNMPDMDG FKLLEHVGLE MDLPVIMMSV DGETSRVMKG VQHGACDYLL 120 KPVRMKELRN IWQHVFRKKI HEIKEVESHD NSSEEIQFLS NEFAERQQML YAGDLNSTLK 180 RKDMTENTDI FVDTDVFDHA SSVKKARVIW TVDLHQKFVD AVNQIGFDKV GPKKILDLMN 240 VPGITRENVA SHLQKYRLYL SRLQKQNYGV TIRGGTTAAT QTDTDGLKDT TNFVLQIQKA 300 VSHSSGKRRN HVDTVTKTEV SGDAALKHAA ISQLQKDSSI KVFGSTSTKT VIPNSQYQIQ 360 MQQQQQHDNH CWNLEGIAAI PFTNYSKHYS NSNQSDYHQH RQQMSTESKQ KHNVSDIKSN 420 HSFSKQINLI SVQVDHGLLG SKNYGIGVSD EFDQQQNYLF DNLDHPQLQK TVPSSSQYEY 480 LNNIDINSMD IHNYNNSTQI TQLQGDMQFD YEFMSMRRM* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 1e-16 | 201 | 263 | 1 | 63 | ARR10-B |
| 5lxu_A | 7e-17 | 205 | 260 | 1 | 56 | Transcription factor LUX |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00010 | PBM | Transfer from AT1G67710 | Download |
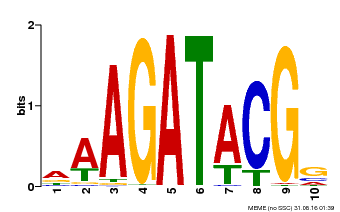
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008797265.1 | 1e-146 | two-component response regulator ORR26-like isoform X1 | ||||
| TrEMBL | A0A0K9P118 | 0.0 | A0A0K9P118_ZOSMR; Two-component response regulator | ||||
| STRING | XP_008797265.1 | 1e-145 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5900 | 37 | 52 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G67710.1 | 1e-110 | response regulator 11 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma4g02000.1 |



