 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma343g00020.1 | ||||||||
| Common Name | ZOSMA_343G00020 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | TCP | ||||||||
| Protein Properties | Length: 435aa MW: 47206.1 Da PI: 4.3649 | ||||||||
| Description | TCP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | TCP | 116.9 | 2.6e-36 | 97 | 200 | 3 | 89 |
TCP 3 gkkdrhskihTkvggRdRRvRlsaecaarfFdLqdeLGfdkdsktieWLlqqakpaikeltgt...............ssssasec..eaess 78
g+kdrhsk++T++g+RdRRvRlsa++a++f+d+qd+LG+d++sk+++WL+++ak+ai+el+++ +ss+a+e+ +++++
Zosma343g00020.1 97 GRKDRHSKVCTAKGPRDRRVRLSAHTAIQFYDVQDRLGYDRPSKAVDWLIKNAKAAIDELAELppwhptattaasgfgPSSPATEMgqDQNDE 189
89*************************************************************999998766666544444443432222222 PP
TCP 79 sssasnsssg......k 89
s s + +
Zosma343g00020.1 190 STS------AafafggS 200
222......12233331 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03634 | 2.3E-30 | 97 | 171 | IPR005333 | Transcription factor, TCP |
| PROSITE profile | PS51369 | 34.086 | 98 | 156 | IPR017887 | Transcription factor TCP subgroup |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009793 | Biological Process | embryo development ending in seed dormancy | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0030154 | Biological Process | cell differentiation | ||||
| GO:0045962 | Biological Process | positive regulation of development, heterochronic | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 435 aa Download sequence Send to blast |
MDDSFLTSCK LEEEGRKKRG SGDPKKPLRP QILAGEEEGE EEEEGNVEEE EIEEDHHRKS 60 SSDQYAVLQR RPVGMRGGHG GGEIVEVEGG HIVRSIGRKD RHSKVCTAKG PRDRRVRLSA 120 HTAIQFYDVQ DRLGYDRPSK AVDWLIKNAK AAIDELAELP PWHPTATTAA SGFGPSSPAT 180 EMGQDQNDES TSAAFAFGGS SSSFLPPSLE SDAIADTIKF FFPIQSSAAS AGERSGGGGG 240 GASIGSSSSA NLAGFHHQQY PHNILSQSQD LRLSLQSFED PILHHQQQQQ QQTCGGPSPE 300 LQSDQSSFFD TPTSVWSEPN QRFFNTGSGY GPYMPALPLH PMFGQNPFFP QRGPLQSNNI 360 NSPSFLALTD PASMATAMQQ ASVMPGDTHG FTSSGFSGFR APARIQGEEE DDDDDDCEYD 420 DDEEDEDEDH QVDI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zkt_A | 4e-20 | 103 | 156 | 1 | 54 | Putative transcription factor PCF6 |
| 5zkt_B | 4e-20 | 103 | 156 | 1 | 54 | Putative transcription factor PCF6 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor playing a pivotal role in the control of morphogenesis of shoot organs by negatively regulating the expression of boundary-specific genes such as CUC genes, probably through the induction of miRNA (e.g. miR164) (PubMed:12931144, PubMed:17307931). Required during early steps of embryogenesis (PubMed:15634699). Participates in ovule develpment (PubMed:25378179). Activates LOX2 expression by binding to the 5'-GGACCA-3' motif found in its promoter (PubMed:18816164). {ECO:0000269|PubMed:12931144, ECO:0000269|PubMed:15634699, ECO:0000269|PubMed:17307931, ECO:0000269|PubMed:18816164, ECO:0000269|PubMed:25378179}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00062 | PBM | Transfer from AT3G15030 | Download |
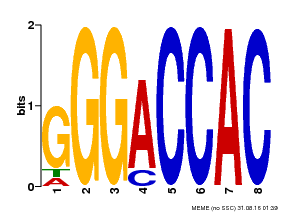
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Repressed by the miRNA miR-JAW/miR319 (PubMed:12931144, PubMed:18816164). {ECO:0000269|PubMed:12931144, ECO:0000269|PubMed:18816164}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010277724.1 | 1e-95 | PREDICTED: transcription factor TCP4 | ||||
| Swissprot | Q8LPR5 | 9e-74 | TCP4_ARATH; Transcription factor TCP4 | ||||
| TrEMBL | A0A0K9P7D7 | 0.0 | A0A0K9P7D7_ZOSMR; TCP family transcription factor TCP4 | ||||
| STRING | XP_010277724.1 | 4e-95 | (Nelumbo nucifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5839 | 33 | 55 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G53230.1 | 1e-45 | TEOSINTE BRANCHED 1, cycloidea and PCF transcription factor 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma343g00020.1 |



