 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma318g00100.1 | ||||||||
| Common Name | ZOSMA_318G00100 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | AP2 | ||||||||
| Protein Properties | Length: 652aa MW: 70938.6 Da PI: 6.6188 | ||||||||
| Description | AP2 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 49 | 1.6e-15 | 308 | 367 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrd.pse.ng..kr.krfslgkfgtaeeAakaaiaarkkleg 55
s+++GV++++++gr++A+++d ++ g ++ ++++lg ++ +e+Aa+a++ a++k++g
Zosma318g00100.1 308 SQFRGVTRHRWTGRYEAHLWDnSCKkEGqtRKgRQVYLGGYDMEEKAARAYDLAALKYWG 367
789******************988886678446*************************98 PP
| |||||||
| 2 | AP2 | 45.7 | 1.7e-14 | 412 | 461 | 3 | 55 |
AP2 3 ykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
y+GV+++++ grW A+I + +k +lg+f t eeAa+a++ a+ k++g
Zosma318g00100.1 412 YRGVTRHHQHGRWQARIGRVAG---NKDLYLGTFSTQEEAAEAYDIAAIKFRG 461
9***************988532...5************************997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 7.85E-16 | 308 | 377 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 2.57E-21 | 308 | 375 | No hit | No description |
| SMART | SM00380 | 5.9E-27 | 309 | 381 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 4.4E-12 | 309 | 367 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 18.769 | 309 | 375 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 2.6E-15 | 309 | 376 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.6E-5 | 310 | 321 | IPR001471 | AP2/ERF domain |
| SuperFamily | SSF54171 | 2.09E-17 | 410 | 470 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 1.30E-24 | 410 | 471 | No hit | No description |
| SMART | SM00380 | 1.2E-33 | 411 | 475 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:3.30.730.10 | 5.2E-18 | 411 | 469 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 19.177 | 411 | 469 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 2.8E-9 | 412 | 461 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 1.6E-5 | 451 | 471 | IPR001471 | AP2/ERF domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007276 | Biological Process | gamete generation | ||||
| GO:0010492 | Biological Process | maintenance of shoot apical meristem identity | ||||
| GO:0042127 | Biological Process | regulation of cell proliferation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 652 aa Download sequence Send to blast |
MSGDQEMKSI MNDNNNSNSN GNWLGFSLSP QMSMEVSADS HPPSEAVNVS SALPNNYFVS 60 PHLNGSGIYY GVEGGNTSGY YSQLSVMPLK SDGSLCIMEA LSRSQQQEGM GPSPSPKLED 120 FLGNSGPTMG SHQYGSNEGE AMALSLDSMY YHHQNPDSLH HHQQHQHNQQ HPYFQSLSEG 180 LCTGLPGHEM CHAPLEGEPM TDDRIPSLKN WVARHYSNGN SGGMGDDGGV GVGSSSVGAM 240 GYGDLQSLSL SMSPGSQSSC VTAPQQILTV ATTDSMVMDN IKKRSNGKSG QKQPVHRKSI 300 DTFGQRTSQF RGVTRHRWTG RYEAHLWDNS CKKEGQTRKG RQVYLGGYDM EEKAARAYDL 360 AALKYWGASS HINFPLENYQ EQLEEMKIMS RQEYVAHLRR KSSGFSRGAS MYRGVTRHHQ 420 HGRWQARIGR VAGNKDLYLG TFSTQEEAAE AYDIAAIKFR GVNAVTNFDI TRYDVEKIAA 480 SSTLLPSDAA RRKNKLLLEA PPQAVVDISN DKNTSRGGGE GGGEEQVQEN TNGNSDWKML 540 LCTQSPPPMQ QSVGYPSGLN GLMSVDMLNS PQVVVDDDSG KIGNTQHMSN ASSLVTSLSS 600 SREDSPDRTS LSMLLSKSSA LPISTWIPST TQMKPAISLA QMAVFAGWTD T* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that recognizes and binds to the DNA consensus sequence 5'-CAC[AG]N[AT]TNCCNANG-3'. Required for the initiation and growth of ovules integumenta, and for the development of female gametophyte. Plays a critical role in the development of gynoecium marginal tissues (e.g. stigma, style and septa), and in the fusion of carpels and of medial ridges leading to ovule primordia. Also involved in organs initiation and development, including floral organs. Maintains the meristematic competence of cells and consequently sustains expression of cell cycle regulators during organogenesis, thus controlling the final size of each organ by controlling their cell number. Regulates INO autoinduction and expression pattern. As ANT promotes petal cell identity and mediates down-regulation of AG in flower whorl 2, it functions as a class A homeotic gene. {ECO:0000269|PubMed:10528263, ECO:0000269|PubMed:10639184, ECO:0000269|PubMed:10948255, ECO:0000269|PubMed:11041883, ECO:0000269|PubMed:12183381, ECO:0000269|PubMed:12271029, ECO:0000269|PubMed:12655002, ECO:0000269|PubMed:8742706, ECO:0000269|PubMed:8742707, ECO:0000269|PubMed:9001406, ECO:0000269|PubMed:9093862, ECO:0000269|PubMed:9118807}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00088 | SELEX | Transfer from AT4G37750 | Download |
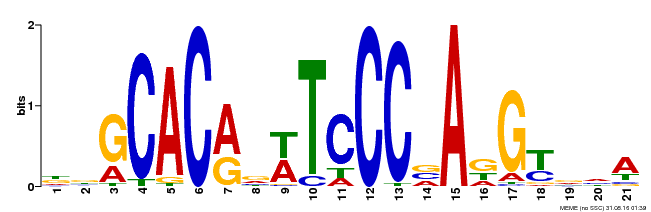
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010939358.1 | 0.0 | AP2-like ethylene-responsive transcription factor CRL5 | ||||
| Swissprot | Q38914 | 1e-155 | ANT_ARATH; AP2-like ethylene-responsive transcription factor ANT | ||||
| TrEMBL | A0A0K9PBF5 | 0.0 | A0A0K9PBF5_ZOSMR; AP2-like ethylene-responsive transcription factor ANT | ||||
| STRING | XP_008785732.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP948 | 38 | 133 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G37750.1 | 1e-122 | AP2 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma318g00100.1 |



