 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma27g00270.1 | ||||||||
| Common Name | ZOSMA_27G00270 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 860aa MW: 96004.5 Da PI: 7.1086 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 74.8 | 9.8e-24 | 165 | 266 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrs 88
f+k+lt sd++++g +++ +++a+e+ g++ + +++l+ +d++g +W++++i+r++++r++l++GW+ Fv++++L +gD ++F + +
Zosma27g00270.1 165 FCKTLTASDTSTHGGFSVLRRHADEClppldmGKQ-PPTQELVAKDLHGVEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL--RGE 255
99*****************************6444.45569************************************************..449 PP
EE..EEEEE-S CS
B3 89 efelvvkvfrk 99
++el+v+v+r+
Zosma27g00270.1 256 NGELRVGVRRA 266
999*****996 PP
| |||||||
| 2 | Auxin_resp | 118.4 | 5.9e-39 | 291 | 373 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+st+++F v+Y+Pr+s+seF+v++++++++lk++ s+GmRfkm+fe+e+++e+r++Gt+vg+sd+d+ rW++SkWrsLk
Zosma27g00270.1 291 AWHAVSTGTMFIVYYKPRTSPSEFIVPYDQYMESLKNNHSIGMRFKMRFEGEEAPEQRFTGTIVGISDADSGRWSGSKWRSLK 373
89*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.9E-40 | 157 | 280 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.26E-38 | 161 | 294 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 2.19E-21 | 163 | 265 | No hit | No description |
| Pfam | PF02362 | 1.4E-21 | 165 | 266 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 11.815 | 165 | 267 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.0E-23 | 165 | 267 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 3.9E-36 | 291 | 373 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 22.873 | 733 | 826 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 1.28E-11 | 735 | 810 | No hit | No description |
| Pfam | PF02309 | 1.4E-6 | 779 | 814 | IPR033389 | AUX/IAA domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009734 | Biological Process | auxin-activated signaling pathway | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 860 aa Download sequence Send to blast |
MTPTSGASQK GNYADGRGES FSSGSSEMND GVKNTVVSDV NRGLSDACPP PVSGKDFDSA 60 FYDELWHACA GPLVTVPKQG EKVYYFPQGH IEQVEASTNQ VANQQMPMYN LQPKILCRVM 120 NVVLKAEPDT DEVFAQITLL PEEDDNTGEK EPPPVVHPRT RVRSFCKTLT ASDTSTHGGF 180 SVLRRHADEC LPPLDMGKQP PTQELVAKDL HGVEWRFRHI FRGQPRRHLL QSGWSVFVSS 240 KRLVAGDAFI FLRGENGELR VGVRRAMKHQ ANVPSSVISS HNMHLGVLAT AWHAVSTGTM 300 FIVYYKPRTS PSEFIVPYDQ YMESLKNNHS IGMRFKMRFE GEEAPEQRFT GTIVGISDAD 360 SGRWSGSKWR SLKVRWDENS SISRPERVSP WKIEPASTPP LNPVPAPRNK RARANHVFPL 420 SNSSSFVTTR DGTKTFVDSH MHEISKVLQG QETKSLHSVF AHDVESEISQ RPTSWIGESK 480 NDGNCQSRLG KTSWKSGARN EPMYSNMLTG FNVSGDTQGL SSVLRRTHED NGPSKNHHQE 540 QDGKFTLLSK PWLMQSNVLD VDRRWESSNQ INVLPNQKID SSRYGGHGYL SLQSYGIDHS 600 TNWSSPKLPP NHIIEGPIKP ILRRPQPCIL KKSELPKVKG SGNCRIFGFQ LNSNPEILDL 660 ETTHINAAHV SDSHSHPASI HQGSGFEIDQ HTDECKGNNS TDSDLVRKEQ VHRLQSSTPT 720 SKDASKPCHG SARSCTKVHK QGIALGRSVD LTKFIDYDQL IDELDLMFDF KGELKPPNKN 780 WVIVYRDNEN DMMLVGDDPW QEFCCIVRKI CIYTPEEVQR MNPGTLSPKI EDGEADISEE 840 RNTATNGTWE SSPSPMADS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-172 | 54 | 398 | 12 | 357 | Auxin response factor 1 |
| 4ldw_A | 1e-172 | 54 | 398 | 12 | 357 | Auxin response factor 1 |
| 4ldw_B | 1e-172 | 54 | 398 | 12 | 357 | Auxin response factor 1 |
| 4ldx_A | 1e-172 | 54 | 398 | 12 | 357 | Auxin response factor 1 |
| 4ldx_B | 1e-172 | 54 | 398 | 12 | 357 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
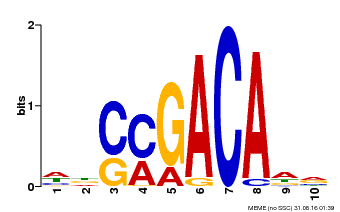
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010942378.1 | 0.0 | auxin response factor 23 | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | A0A0K9PFI0 | 0.0 | A0A0K9PFI0_ZOSMR; Auxin response factor | ||||
| STRING | XP_010249208.1 | 0.0 | (Nelumbo nucifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1178 | 36 | 111 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma27g00270.1 |



