 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Zosma1g03840.1 | ||||||||
| Common Name | ZOSMA_1G03840 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Alismatales; Zosteraceae; Zostera
|
||||||||
| Family | YABBY | ||||||||
| Protein Properties | Length: 261aa MW: 29648.4 Da PI: 10.6509 | ||||||||
| Description | YABBY family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | YABBY | 108 | 1.8e-33 | 98 | 237 | 7 | 162 |
YABBY 7 seqvCyvqCnfCntilavsvPstslfkvvtvrCGhCtsllsvnlakasqllaaeshldeslkeelleelkveeenlksnvekeesastsvssekl 101
++ +C + + t+++v vP + ++++vtv+CGhC+++ ++ ++ + + ++ + +l+ +++ + ++ ++s+ +ss
Zosma1g03840.1 98 PQSICAMFVATSATLFLVGVPLKRIMETVTVKCGHCSNISFLT------SRPIMP----TNLPDFHMTLGLQTSCH---DFRKGQPSSPTSS--- 176
678999999999*************************985532......222222....34566777777665544...4445555555555... PP
YABBY 102 senedeevprvppvirPPekrqrvPsaynrfikeeiqrikasnPdishreafsaaaknWah 162
en e +p+v++v++PPe++ rvPsaynrf+keeiqrik++ P+i hreafs+aaknWa
Zosma1g03840.1 177 VENLAEMTPKVHHVVKPPERKHRVPSAYNRFMKEEIQRIKTAQPNIPHREAFSMAAKNWAK 237
346778899999***********************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF04690 | 1.2E-38 | 99 | 237 | IPR006780 | YABBY protein |
| SuperFamily | SSF47095 | 5.11E-8 | 185 | 238 | IPR009071 | High mobility group box domain |
| Gene3D | G3DSA:1.10.30.10 | 1.2E-4 | 191 | 237 | IPR009071 | High mobility group box domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010254 | Biological Process | nectary development | ||||
| GO:0010582 | Biological Process | floral meristem determinacy | ||||
| GO:0048479 | Biological Process | style development | ||||
| GO:0016021 | Cellular Component | integral component of membrane | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 261 aa Download sequence Send to blast |
MRLVLFLYVW YVRVFFLLLN LSVRKRWRCV VDEKGSLYKS GGRRDSEGDE WKQLKQLRKK 60 QDITRCEHKY VFKASLFKAS YHQTATQFSS SSFSSWIPQS ICAMFVATSA TLFLVGVPLK 120 RIMETVTVKC GHCSNISFLT SRPIMPTNLP DFHMTLGLQT SCHDFRKGQP SSPTSSVENL 180 AEMTPKVHHV VKPPERKHRV PSAYNRFMKE EIQRIKTAQP NIPHREAFSM AAKNWAKSDT 240 NAPNGSQKSP KAAAAISLHV * |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Regulates carpel specification in flower development. Severe or intermediate mutation in DL causes complete or partial homeotic conversion of carpels into stamens without affecting the identities of other floral organs. Interacts antagonistically with class B genes and controls floral meristem determinacy. Regulates midrib formation in leaves probably by inducing cell proliferation in the central region of the leaf. {ECO:0000269|PubMed:14729915}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00220 | DAP | Transfer from AT1G69180 | Download |
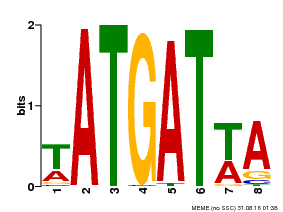
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010936439.1 | 1e-51 | protein DROOPING LEAF | ||||
| Swissprot | Q76EJ0 | 7e-48 | YABDL_ORYSJ; Protein DROOPING LEAF | ||||
| TrEMBL | A0A0K9PN40 | 0.0 | A0A0K9PN40_ZOSMR; Protein DROOPING LEAF | ||||
| STRING | TRIUR3_31693-P1 | 4e-48 | (Triticum urartu) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5951 | 36 | 57 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69180.1 | 2e-42 | YABBY family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Zosma1g03840.1 |



